Patient-Derived Xenografts vs. Organoids: A Modern Guide to Translational Cancer Models for Research & Drug Discovery
This comprehensive guide explores the critical role of Patient-Derived Xenografts (PDXs) and Organoids (PDOs) in modern translational cancer research.
Patient-Derived Xenografts vs. Organoids: A Modern Guide to Translational Cancer Models for Research & Drug Discovery
Abstract
This comprehensive guide explores the critical role of Patient-Derived Xenografts (PDXs) and Organoids (PDOs) in modern translational cancer research. Targeted at researchers, scientists, and drug development professionals, it provides a foundational understanding of both models, detailing their methodologies, applications in preclinical studies, and use in personalized medicine. The article delves into practical challenges, offering troubleshooting and optimization strategies for model establishment and fidelity. A comparative analysis validates the complementary strengths and limitations of PDXs and PDOs for specific research intents. By synthesizing current best practices, this resource aims to empower the selection and implementation of these powerful tools to accelerate oncology research and therapeutic development.
What Are PDX and PDO Models? Core Concepts and Evolution in Cancer Biology
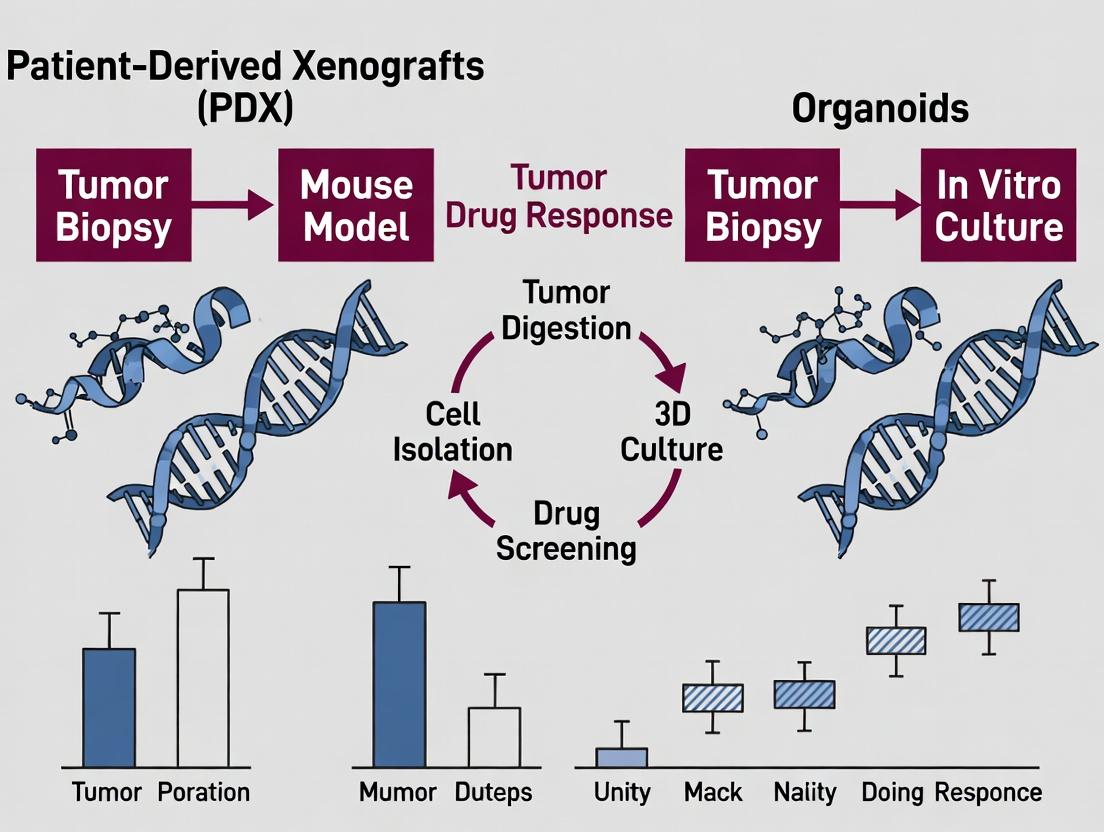
Within the modern oncology research paradigm, patient-derived xenografts (PDXs) and patient-derived organoids (PDOs) represent two pillars of translational cancer research. This whitepaper focuses on PDXs, which are established by the direct transplantation of freshly resected human tumor tissue into immunodeficient animal hosts. PDXs serve as a critical preclinical model system that retains the genetic heterogeneity, histopathology, and molecular characteristics of the original patient tumor. They are indispensable for studying tumor biology, metastasis, and for the in vivo evaluation of therapeutic efficacy, often complementing data generated from organoid cultures.
Key Methodological Workflow and Protocols
The generation and utilization of PDXs follow a standardized, multi-stage workflow. Success hinges on careful sample acquisition, host selection, and subsequent passaging.
2.1 Core Experimental Protocol: PDX Generation and Expansion
- Sample Acquisition & Processing: Fresh tumor tissue from surgical resection or biopsy is collected in sterile, cold transport medium (e.g., DMEM/F12 with 10% FBS and 1% penicillin/streptomycin). Tissue is mechanically dissociated and/or enzymatically digested (e.g., with collagenase/hyaluronidase mix) into small fragments (approx. 1-3 mm³).
- Host Animal & Transplantation: Immunodeficient mice (e.g., NOD-scid IL2Rγnull (NSG)) are the gold standard host. Fragments are implanted subcutaneously (most common), orthotopically (into the organ/tissue of origin), or via other routes using a trocar. Animals are monitored for tumor engraftment.
- Engraftment & Passaging: Successful engraftment is defined by tumor growth to a predetermined volume (e.g., 500-1500 mm³). The primary xenograft (P0 or F1) is then harvested, divided, and re-implanted into new host animals to create subsequent passages (F2, F3, etc.). Cryopreservation of tumor fragments at each passage is essential for biobanking.
2.2 PDX Experimentation Workflow Diagram
Title: PDX Generation and Experimental Workflow
Quantitative Data on PDX Characteristics
Table 1: PDX Engraftment Rates and Stability Across Major Cancer Types
| Cancer Type | Typical Engraftment Rate Range (%) | Median Latency to Growth (Weeks) | Reported Genomic Stability (Passages) | Key Considerations |
|---|---|---|---|---|
| Pancreatic Ductal Adenocarcinoma | 50 - 75 | 8 - 16 | Stable for >10 passages | Stromal replacement by murine cells occurs early. |
| Triple-Negative Breast Cancer | 60 - 80 | 6 - 12 | Stable for 5-8 passages | Higher engraftment vs. hormone receptor-positive subtypes. |
| Colorectal Carcinoma | 65 - 85 | 4 - 10 | Stable for >10 passages | Often retains original mutational profile and drug response. |
| Non-Small Cell Lung Cancer | 30 - 50 | 10 - 20 | Stable for 4-6 passages | Lower engraftment; can be influenced by driver mutations. |
| Glioblastoma | 40 - 60 | 12 - 24 | Variable | Often requires orthotopic implantation for relevant biology. |
| Melanoma | 70 - 90 | 4 - 8 | Stable for 5-10 passages | High success rate; can be used for metastasis studies. |
Table 2: Comparison of Common Immunodeficient Mouse Strains for PDX
| Mouse Strain | Immune Deficiency Characteristics | Key Advantages for PDX | Key Limitations |
|---|---|---|---|
| NOD-scid IL2Rγnull (NSG) | Lack T, B, NK cells; deficient in cytokine signaling. | Highest engraftment rates; supports hematopoietic malignancies. | Extremely immunocompromised; requires strict SPF conditions. |
| NOD-scid | Lack T and B cells; functional NK cells and macrophages. | More robust health than NSG; good for many solid tumors. | Lower engraftment than NSG due to innate immunity. |
| BALB/c nude | Athymic; lack T cells, have B and NK cells. | Good general health and lifespan. | Lower engraftment; not suitable for lymphoid tumors. |
| C.B-17 scid | Lack T and B cells; functional NK cells and complement. | Historical standard. | "Leaky" immunity with age; lower engraftment than NSG. |
Signaling Pathways in PDX Tumor-Stroma Interactions
A critical phenomenon in PDX models is the interaction between human tumor cells and the murine stromal microenvironment, which evolves upon passaging.
Title: Key Signaling in PDX Human-Murine Crosstalk
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Materials for PDX Research
| Item/Category | Specific Example(s) | Function & Application |
|---|---|---|
| Transport/Storage Medium | DMEM/F12 + 10% FBS + 1% P/S; Tissue Storage Solutions (e.g., commercial cryopreservation media). | Maintains tissue viability from OR to lab; enables cryopreservation of tumor fragments for biobanking. |
| Tissue Dissociation Kits | GentleMACS Dissociator with human Tumor Dissociation Kits; Collagenase/Hyaluronidase mixes. | Generates single-cell suspensions or small fragments for implantation or downstream analysis (e.g., flow cytometry). |
| Matrigel / ECM | Growth Factor Reduced Matrigel. | Mixed with tumor fragments for implantation to enhance engraftment, particularly for low-volume samples. |
| Immunodeficient Mice | NOD-scid IL2Rγnull (NSG) mice. | Gold-standard host organism with profound immunodeficiency enabling high engraftment of human tissues. |
| Species-Specific Antibodies | Anti-human HLA-ABC; Anti-mouse H-2Kd; Anti-human/mouse CD31 (dual-species staining). | Critical for IHC/IF to distinguish human tumor cells from infiltrating murine stroma and vasculature. |
| Genomic Analysis Kits | Human-specific STR Profiling Kits; Mouse Contamination Detection Assays; ddPCR assays with human-specific probes. | Authenticates PDX model (matches to patient), monitors for cross-contamination, and quantifies human DNA. |
| In Vivo Imaging Reagents | Luciferin (for bioluminescence); Species-specific antibody-conjugated fluorescent probes. | Enables non-invasive tracking of tumor growth and metastasis, and visualization of human vs. murine components. |
This whitepaper serves as a technical guide within a broader thesis exploring patient-derived xenografts (PDXs) and organoids in cancer research. While PDXs involve implanting human tumor tissue into immunodeficient mice to create in vivo models, Patient-Derived Organoids (PDOs) represent a complementary in vitro paradigm. PDOs are three-dimensional, self-organizing microtissues derived from patient stem cells or tumor fragments that recapitulate the architecture, heterogeneity, and genetic landscape of the original tissue. Their development addresses key limitations of PDXs, including cost, throughput, and scalability for drug screening and functional precision oncology.
Core Principles and Current Data
PDOs are generated from either adult stem cells (ASCs) or induced pluripotent stem cells (iPSCs) cultured in a basement membrane matrix (e.g., Matrigel) and fed with a tailored cocktail of growth factors that mimic the native stem cell niche. This environment supports the expansion and self-organization of cell populations into structures exhibiting key functional properties of the organ of origin.
Table 1: Comparative Metrics of PDOs vs. Traditional Models in Cancer Research
| Metric | Patient-Derived Organoids (PDOs) | Patient-Derived Xenografts (PDXs) | 2D Cell Lines |
|---|---|---|---|
| Establishment Success Rate | ~30-80% (varies by cancer type) | ~20-40% | N/A (Immortalized) |
| Establishment Time | 2-8 weeks | 3-6 months | N/A |
| Genetic Stability | High (> 6 months in culture) | High, but mouse stroma invasion | Low (cultured long-term) |
| Throughput for Drug Screening | High (96-/384-well formats) | Very Low (expensive, low scalability) | High |
| Cost per Model/Line | Low-Medium | Very High | Very Low |
| Tumor Microenvironment | Limited (mostly epithelial) | Preserved (human tumor in mouse stroma) | Absent |
| Clinical Predictive Value (Ongoing) | High in retrospective studies | Historically high | Low |
Data synthesized from recent (2023-2024) reviews and studies in *Nature Protocols, Cell, and Cancer Discovery.*
Detailed Experimental Protocol: Generating Colorectal Cancer PDOs
Title: Protocol for Establishing and Maintaining Colorectal Cancer Patient-Derived Organoids.
Materials:
- Fresh colorectal cancer biopsy or surgical specimen (in cold Advanced DMEM/F12 + antibiotics).
- Digestion Solution: Advanced DMEM/F12, 1 mg/ml Collagenase II, 10 µM Y-27632 (ROCK inhibitor).
- Basement Membrane Extract: Matrigel, Growth Factor Reduced, Phenol Red-free.
- Complete Human Intestinal Organoid Medium:
- Basal: Advanced DMEM/F12, 10 mM HEPES, 1x GlutaMAX.
- Additives: 1x B-27 Supplement, 1x N-2 Supplement.
- Growth Factors: 50 ng/ml human EGF, 100 ng/ml human Noggin, 500 ng/ml human R-spondin-1 (crucial for Wnt pathway activation), 10 nM Gastrin I, 1 mM N-acetylcysteine, 10 mM Nicotinamide.
- Antibiotics: Primocin (100 µg/ml).
- ROCK Inhibitor: 10 µM Y-27632 (for first 2-3 days post-seeding).
Methodology:
- Tissue Processing: Mince tissue into <1 mm³ fragments. Wash with cold basal medium.
- Enzymatic Digestion: Incubate fragments in digestion solution at 37°C for 30-60 mins with gentle agitation. Pipette vigorously every 15 mins.
- Filtration & Washing: Pass digest through a 100 µm cell strainer. Centrifuge filtrate at 300 x g for 5 mins. Wash pellet with basal medium.
- Embedding in Matrix: Resuspend cell pellet in cold BME/Matrigel (~50-100 µl per dome). Plate 10-20 µl domes in a pre-warmed 48-well plate. Polymerize at 37°C for 20-30 mins.
- Culture Initiation: Overlay each dome with 300 µl of complete medium + Y-27632. Culture at 37°C, 5% CO₂.
- Maintenance & Passaging: Change medium every 2-3 days. Monitor for organoid formation (typically within 3-7 days). Passage every 7-14 days by mechanically breaking organoids and re-embedding in fresh Matrigel.
Key Signaling Pathways in PDO Growth and Maintenance
Diagram 1: Wnt/β-catenin Signaling Niche in Intestinal PDOs (86 chars)
Experimental Workflow for PDO-Based Drug Screening
Diagram 2: PDO High-Throughput Drug Screening Pipeline (75 chars)
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for PDO Culture and Assays
| Reagent Category | Specific Product/Component | Function in PDO Research |
|---|---|---|
| Basement Matrix | Corning Matrigel, GFR, Phenol Red-free; Cultrex BME | Provides a 3D scaffold mimicking the extracellular matrix for organoid growth and polarization. |
| Niche Factor Cocktail | Recombinant Human R-spondin-1, Noggin, EGF, Wnt-3a | Essential growth factors that maintain stemness and drive proliferation in epithelial organoids. |
| Base Medium | Advanced DMEM/F-12 | A stable, low-osmolarity basal medium optimized for organoid culture. |
| Serum-Free Supplements | B-27 Supplement (50x), N-2 Supplement (100x) | Provide hormones, proteins, and essential nutrients for survival and growth in defined media. |
| Dissociation Agent | Accutase; Collagenase/Dispase | Enzymatic solutions for breaking down organoids into single cells or small clusters for passaging or seeding assays. |
| ROCK Inhibitor | Y-27632 dihydrochloride | Enhances survival of single stem cells by inhibiting apoptosis during passaging and seeding. |
| Viability Assay | CellTiter-Glo 3D | Luminescent ATP-based assay optimized for 3D cultures to measure cell viability for drug screening. |
| Cryopreservation Medium | CryoStor CS10 | A serum-free, DMSO-containing solution for freezing and long-term storage of PDO biobanks. |
The study of cancer has undergone a paradigm shift, moving from simplistic two-dimensional (2D) cell line models to sophisticated, patient-specific three-dimensional (3D) systems. This evolution is central to the broader thesis on Introduction to patient-derived xenografts and organoids in cancer research, which posits that increasing biological fidelity is paramount for translational relevance. Traditional immortalized cancer cell lines, while invaluable for foundational discovery, often fail to recapitulate the genetic heterogeneity, cellular architecture, and drug response observed in human tumors. This historical progression—from cell lines to patient-derived xenografts (PDXs) and patient-derived organoids (PDOs)—represents a concerted effort to bridge the gap between bench findings and clinical success.
Quantitative Comparison of Model Systems
The table below summarizes key quantitative metrics that distinguish these model systems, highlighting the trade-offs between throughput, fidelity, and clinical correlation.
Table 1: Quantitative Comparison of Preclinical Cancer Models
| Feature | Immortalized 2D Cell Lines | Patient-Derived Xenografts (PDXs) | Patient-Derived Organoids (PDOs) |
|---|---|---|---|
| Establishment Success Rate | ~100% (from existing banks) | 20-40% (varies by cancer type) | 50-80% (varies by cancer type) |
| Time to Usable Model | Immediate (from bank) | 4-8 months (engraftment & expansion) | 2-4 weeks (from biopsy) |
| Genetic Drift | High (long-term culture) | Low (early passages) | Very Low (limited in vitro culture) |
| Stromal/Immune Component | Absent or minimal | Human tumor, murine stroma/immune system | Human tumor epithelium only |
| Throughput (Drug Screens) | Very High (96/384-well plates) | Low (cost/time intensive) | Medium-High (3D formats possible) |
| Clinical Predictive Value (PPV/NPV*) | Low (~5-10%) | High for some tumor types (~80-90%) | Emerging, promising (~70-85%) |
| Cost per Model | Low ($100s) | Very High ($10,000s) | Medium ($1,000s) |
*PPV: Positive Predictive Value; NPV: Negative Predictive Value for drug response.
Detailed Experimental Protocols
Protocol: Generation of Patient-Derived Tumor Organoids (PDOs)
Objective: To establish a 3D in vitro culture system that retains the genetic and phenotypic characteristics of a patient's tumor.
Materials: See "The Scientist's Toolkit" section below.
Methodology:
- Tissue Processing: Mince fresh tumor biopsy or surgical specimen (1-5 mm³) in cold Advanced DMEM/F12 using scalpels. Digest with 5 mL of enzymatic mix (Collagenase II, Dispase, DNase I) for 30-60 minutes at 37°C with agitation.
- Cell Isolation: Pass the digest through a 70 µm cell strainer. Wash the flow-through with cold buffer and centrifuge at 300 x g for 5 minutes.
- Matrix Embedding: Resuspend the pellet in cold, growth factor-reduced Basement Membrane Extract (BME). Plate 30 µL droplets (~10,000 cells/droplet) in a pre-warmed 24-well plate. Polymerize for 30-45 minutes at 37°C.
- Culture: Overlay each dome with 500 µL of complete organoid growth medium, tailored to the cancer of origin (e.g., IntestiCult for CRC, proprietary mixes for pancreas/breast). Culture at 37°C, 5% CO₂.
- Passaging: Every 7-14 days, mechanically disrupt BME domes, recover organoids, and dissociate with TrypLE Express for 5-10 minutes. Re-embed fragments or single cells in fresh BME.
Protocol: Establishment of Patient-Derived Xenografts (PDXs)
Objective: To engraft and propagate patient tumor tissue in an immunocompromised mouse host, preserving tumor histopathology and heterogeneity.
Methodology:
- Host Preparation: Use 6-8 week-old NOD-scid-IL2Rγ[null] (NSG) or similar mice. House in specific pathogen-free (SPF) conditions.
- Implantation: Process fresh tumor tissue as in Step 3.1, but do not fully dissociate. Create 1-2 mm³ fragments in sterile PBS.
- Surgical Implantation (Subrenal Capsule): Anesthetize mouse. Make a small flank incision, expose kidney, and place one tumor fragment under the renal capsule using forceps. Close wound.
- Alternative Implantation (Subcutaneous): Mix tumor fragments 1:1 with BME and inject 100 µL subcutaneously into the flank.
- Monitoring: Measure tumor volume twice weekly using calipers (Volume = (Length x Width²)/2). Endpoint is typically at 1500 mm³.
- Passaging: Excise the xenograft, necrotic tissue removed, and re-implant fragments into a new cohort of mice (P1 passage). Early passages (P1-P3) are used for experiments.
Visualizing Workflows and Pathways
Title: Evolution from Patient to Preclinical Models
Title: Key Oncogenic Signaling Pathways in Cancer
The Scientist's Toolkit: Essential Research Reagents
Table 2: Key Reagent Solutions for Patient-Derived Models
| Reagent/Material | Function | Example Product/Brand |
|---|---|---|
| Basement Membrane Extract (BME) | Provides a 3D scaffold mimicking the extracellular matrix; essential for organoid growth and polarization. | Corning Matrigel, Cultrex BME |
| Advanced DMEM/F12 | Base medium for organoid culture, optimized for low serum/no serum conditions. | Thermo Fisher Scientific |
| R-spondin 1 Conditioned Media | Activates Wnt signaling, critical for stem cell maintenance in gastrointestinal organoids. | Produced in-house from RSPO1-293T cells or commercial (e.g., PeproTech). |
| Noggin Conditioned Media | BMP pathway inhibitor; essential for maintaining stemness in epithelial organoids. | Produced in-house from Noggin-293T cells. |
| Y-27632 (ROCK Inhibitor) | Inhibits anoikis (cell death due to detachment); increases survival of dissociated single cells during seeding/passaging. | STEMCELL Technologies (RevitaCell) |
| Collagenase/Dispase Enzymes | Enzymatic cocktail for digesting solid tumor tissue into small cell aggregates/clusters. | STEMCELL Technologies (Gentle Cell Dissociation Reagent) |
| NSG Mice | Immunocompromised host for PDX studies; lacks T, B, and NK cells, enabling high engraftment rates of human tissue. | The Jackson Laboratory (NOD.Cg-Prkdc |
| TrypLE Express | Gentle, animal-origin-free recombinant protease for dissociating organoids into single cells or small clusters. | Thermo Fisher Scientific |
Within the broader thesis on Introduction to patient-derived xenografts (PDXs) and organoids in cancer research, this guide addresses the foundational pillar of preserving native tumor architecture. The translational validity of PDX and organoid models is fundamentally dependent on their fidelity to the original patient tumor, encompassing both the diverse clonal populations of cancer cells (heterogeneity) and the complex milieu of stromal, immune, and vascular components (microenvironment). This principle is paramount for ensuring research outcomes in drug screening, biomarker discovery, and personalized medicine are clinically relevant.
The Imperative of Preservation: Data-Driven Rationale
The failure to maintain heterogeneity and the TME leads to model drift and loss of predictive power. Key quantitative findings underscore this necessity.
Table 1: Impact of Preservation on Model Fidelity and Clinical Correlation
| Metric | Low-Preservation Model (e.g., cell line) | High-Preservation Model (PDX/Organoid) | Clinical Correlation Evidence |
|---|---|---|---|
| Genetic Heterogeneity | Low (clonal dominance) | High (maintains subclonal diversity) | PDX models retain ~80-95% of somatic mutations from donor tumor across early passages (1-4). |
| Stromal Cell Content | Near zero | Variable, can be sustained | Early passage PDXs retain human stromal cells (typically <5% by passage 4), replaced by murine stroma thereafter. |
| Drug Response Predictivity | Moderate (40-50%) | High (70-90%) | A 2022 cohort study showed PDX drug response predicted clinical outcome with 87% accuracy in NSCLC. |
| Transcriptomic Stability | High drift from origin | Stable for critical passages | Organoids show >90% transcriptome concordance with parent tumor for at least 6 months in culture. |
Detailed Methodological Protocols
Protocol 1: Minimally Biased Processing for PDX Generation
Objective: To engraft tumor tissue into immunodeficient mice while minimizing selection pressure on tumor cell populations.
- Tissue Collection: Obtain fresh tumor tissue in cold, serum-free, antibiotic-containing transport medium (e.g., DMEM/F12) within 1 hour of resection/biopsy.
- Mechanical Dissociation: Mince tissue into ~1-3 mm³ fragments using sterile scalpels. Avoid enzymatic digestion where possible to preserve cell-cell interactions and viability of non-epithelial cells.
- Implantation:
- For subcutaneous engraftment, mix 2-3 fragments with 50µL of Matrigel (basement membrane matrix) and implant into the flanks of NOD-scid IL2Rγ[null] (NSG) mice.
- For orthotopic engraftment, implant a single fragment directly into the cognate organ (e.g., liver, breast fat pad) of anesthetized mice.
- Passaging & Biobanking: Upon tumor reach 1.5 cm³, harvest, and repeat Step 2. A portion is cryopreserved in 90% FBS/10% DMSO, and the rest is passaged. Limit passages to <5 for most studies to retain TME characteristics.
Protocol 2: Establishing Patient-Derived Organoid (PDO) Co-Cultures
Objective: To establish in vitro 3D organoids that include critical TME components, such as cancer-associated fibroblasts (CAFs).
- Enzymatic Dissociation: Digest minced tumor tissue for 30-60 mins at 37°C in an enzyme cocktail (e.g., Collagenase/Hyaluronidase/DNase I) with gentle agitation.
- Stromal Cell Enrichment: Filter digestate through a 100µm strainer. The flow-through contains single cells enriched for immune and stromal cells. Pellet by centrifugation (300 x g, 5 min).
- Epithelial Organoid Culture: The larger fragments retained on the strainer are embedded in 50µL BME (Basement Membrane Extract) domes in 24-well plates. Overlay with organoid-specific medium (e.g., Advanced DMEM/F12 supplemented with Noggin, R-spondin, WNT agonists, and tissue-specific growth factors).
- CAF Co-Culture: Plate the stromal-enriched single-cell pellet in a separate well in fibroblast medium (DMEM + 10% FBS) to establish a CAF monolayer. After 3-5 passages, seed CAFs around the BME dome containing the epithelial organoids to establish a paracrine signaling co-culture system.
Visualization of Key Concepts
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for Preserving Heterogeneity & TME
| Item | Function & Rationale |
|---|---|
| NSG (NOD-scid IL2Rγ[null]) Mice | Gold-standard immunodeficient host for PDX; lacks T, B, and NK cells, enabling high engraftment rates of human tumors with residual human immune components in early passages. |
| Basement Membrane Extract (BME) / Matrigel | Solubilized basement membrane proteins that polymerize at 37°C to provide a 3D scaffold for organoid growth, mimicking the in vivo ECM and supporting polarized epithelial structures. |
| R-spondin-1 & Noggin | Critical growth factor supplements for organoid media. Inhibit differentiation and promote stemness/self-renewal of epithelial cells across many tissues, enabling long-term culture. |
| Collagenase/Hyaluronidase | Enzyme blend for gentle dissociation of tumor tissue, preserving cell surface receptors and viability better than trypsin, crucial for maintaining stromal cell populations. |
| Fetal Bovine Serum (FBS), Charcoal-Stripped | For CAF and stromal cell culture. Charcoal-stripping removes hormones and growth factors, reducing variability and unwanted stimulation in co-culture experiments. |
| Cryopreservation Medium (90% FBS/10% DMSO) | For long-term biobanking of tumor fragments and early-passage organoids. Slow freezing protocols are essential to maintain viability of all cell types upon thaw. |
| Human-Specific Flow Cytometry Antibodies (e.g., anti-hCD45, hEpCAM, hVimentin) | To quantify and isolate human vs. murine cells in PDX models and characterize epithelial/stromal/immune compartments in both PDX and organoids. |
The study of cancer biology and the development of novel therapeutics have long relied on traditional two-dimensional (2D) cell culture. While this method has provided foundational knowledge, its limitations are increasingly apparent within the modern paradigm of precision oncology. This guide frames the critical advantages of advanced three-dimensional (3D) models—specifically patient-derived xenografts (PDXs) and organoids—within the broader thesis of their indispensable role in cancer research. These systems bridge the gap between simplistic monocultures and the immense complexity of human tumors, offering a more physiologically relevant platform for mechanistic studies, drug screening, and personalized medicine strategies.
Quantitative Limitations of 2D Culture: A Data-Driven Critique
The following table summarizes key quantitative discrepancies between 2D culture and native tumor biology, highlighting the rationale for adopting advanced models.
Table 1: Comparative Analysis of 2D Culture vs. In Vivo Tumor Physiology
| Biological Feature | Traditional 2D Culture | In Vivo Tumor Microenvironment | Impact on Research Fidelity |
|---|---|---|---|
| Proliferation Gradients | Homogeneous, high proliferation. | Heterogeneous (hypoxic, nutrient-deprived cores). | Overestimates drug efficacy; fails to model quiescent cell populations. |
| Gene Expression Profiles | Altered significantly within days in vitro; loss of native signatures. | Maintains patient- and region-specific expression patterns. | Poor predictor of clinical tumor behavior and drug response. |
| Drug Response IC50 Values | Often 10-1000x lower than in vivo results. | Aligns with clinical dosing ranges in PDX/organoid models. | Leads to false positives in preclinical screens. |
| Stromal Composition | Lacks critical stromal cells (CAFs, immune cells, endothelial cells). | Complex stroma contributing to signaling, immunity, and drug resistance. | Neglects stroma-mediated resistance and therapeutic targets. |
| Metabolic State | Primarily glycolytic, driven by high oxygen and glucose. | Heterogeneous: oxidative phosphorylation, glycolysis, and fatty acid oxidation. | Misrepresents metabolic vulnerabilities and drug metabolism. |
Core Advantages of 3D Patient-Derived Models
Architectural and Mechanical Cues
3D cultures recapitulate cell-cell and cell-extracellular matrix (ECM) interactions, restoring apical-basal polarity, tissue structure, and mechanical signaling. This architecture directly influences critical oncogenic pathways.
Genetic and Phenotypic Stability
PDXs and organoids maintain the genetic, transcriptomic, and histopathological features of the patient tumor of origin over prolonged periods, unlike 2D lines which undergo rapid clonal selection and genetic drift.
Predictive Validity for Drug Response
Multiple studies have demonstrated that PDX and organoid models show superior correlation with patient clinical response compared to 2D lines. The table below summarizes representative data.
Table 2: Predictive Performance of Advanced Models vs. 2D in Drug Screening
| Study (Example) | Model Type | Correlation with Patient Response | 2D Line Correlation | Key Finding |
|---|---|---|---|---|
| Gao et al., 2015 | PDX Panel (n>1000) | ~90% positive predictive value (PPV) for phase II outcomes. | Not directly comparable; historically low. | PDX "avatars" recapitulate treatment responses seen in donors. |
| Vlachogiannis et al., 2018 | PDOs (GI Cancers) | 100% PPV, 93% negative predictive value (NPV). | N/A | Organoid pharmacotyping predicted clinical outcome in interventional trial. |
| Ooft et al., 2019 | PDOs (Metastatic CRC) | 80% sensitivity, 100% specificity. | <50% sensitivity in historical data. | Guided patient stratification for irinotecan-based chemotherapy. |
Experimental Protocols for Key Applications
Protocol: Establishing a Patient-Derived Organoid (PDO) Biobank for Drug Screening
Objective: To generate, expand, and cryopreserve living organoids from fresh tumor tissue for high-throughput pharmacotyping.
Materials: See "The Scientist's Toolkit" below. Workflow:
- Tissue Processing: Mince fresh tumor biopsy (<1 hour from resection) into <1 mm³ fragments in cold Advanced DMEM/F12. Digest with collagenase/hyaluronidase for 30-120 mins at 37°C.
- Cell Isolation: Filter through 100μm strainer. Centrifuge at 300-500 x g for 5 min. Lyse red blood cells if needed. Wash in basal medium.
- Embedding & Seeding: Resuspend cell pellet in Matrigel (or BME) on ice. Plate 30-50 μL domes in pre-warmed 24-well plate. Polymerize for 20-30 min at 37°C.
- Culture: Overlay with complete organoid growth medium, tailored to cancer type (e.g., containing Noggin, R-spondin, EGF, Wnt3a for GI cancers). Replace medium every 2-3 days.
- Passaging: Mechanically and enzymatically disrupt organoids (TrypLE) every 7-14 days. Re-embed fragments in fresh Matrigel.
- Cryopreservation: Harvest organoids, resuspend in freezing medium (90% FBS, 10% DMSO), cool at -1°C/min in isopropanol chamber, store in liquid nitrogen.
- Drug Assay: Dissociate to single cells or small clusters, seed in Matrigel, treat with compound library after 3-5 days. Assess viability via CellTiter-Glo 3D after 5-7 days.
Protocol:In VivoDrug Efficacy Study Using PDX Models
Objective: To evaluate the antitumor activity of a novel compound in a PDX model with known genetic background.
Materials: Immunocompromised mice (NSG, NOG), PDX tumor fragment, calipers, dosing agents. Workflow:
- Tumor Implantation: Subcutaneously implant a 30-60 mm³ PDX tumor fragment (from passage mouse) into the flank of anesthetized host mouse using a trocar.
- Randomization: When tumors reach 100-200 mm³, randomize mice into vehicle control and treatment groups (n=5-10/group).
- Dosing: Administer compound or vehicle via predetermined route (oral, IP, IV) at established schedule (e.g., QDx21).
- Monitoring: Measure tumor volume (0.5 x length x width²) and body weight 2-3 times weekly.
- Endpoint Analysis: At study end, euthanize mice. Harvest tumors for weight measurement, formalin fixation (for IHC), and snap-freezing (for molecular analysis). Calculate %TGI (Tumor Growth Inhibition) vs. control.
- Statistical Analysis: Compare tumor volumes/growth curves between groups using repeated measures ANOVA.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents for Advanced 3D Cancer Models
| Reagent/Material | Function & Role | Example Product/Brand |
|---|---|---|
| Basement Membrane Extract (BME) | Provides a 3D scaffold with physiological ECM proteins (laminin, collagen IV) for organoid growth and polarization. | Corning Matrigel, Cultrex BME |
| Tissue Dissociation Enzymes | Gentle digestion of tumor tissue to single cells or small clusters while preserving viability and stemness. | Collagenase/Hyaluronidase blend, Liberase, TrypLE Express |
| Defined Growth Factor Cocktails | Supplements to mimic specific stem cell niches (e.g., Wnt, R-spondin, Noggin for intestinal crypt). | Recombinant human EGF, Wnt3a, R-spondin-1, Noggin |
| Rho-Kinase (ROCK) Inhibitor | Enhances survival of single cells and stem cells post-dissociation by inhibiting anoikis. | Y-27632 (dihydrochloride) |
| 3D-Optimized Viability Assay | Luciferase-based ATP quantification reagent engineered to penetrate 3D structures. | CellTiter-Glo 3D (Promega) |
| Immunocompromised Mouse Strains | Host for PDX engraftment, lacking adaptive immunity to tolerate human tissue. | NOD-scid IL2Rγnull (NSG), NOG mice |
How to Establish and Use PDX & PDO Models: Protocols and Preclinical Applications
Patient-derived xenografts (PDXs) are a cornerstone of modern translational oncology, providing a model system that more faithfully recapitulates tumor heterogeneity, architecture, and drug response compared to traditional cell lines. As part of a comprehensive exploration of Introduction to patient-derived xenografts and organoids in cancer research, this guide details the critical hands-on protocols for establishing and maintaining PDX lines. This in vivo approach, complemented by in vitro organoid models, provides a powerful platform for studying tumor biology and preclinical drug evaluation.
Pre-Implantation: Tumor Tissue Processing
Objective: To prepare viable tumor tissue from a patient (or donor mouse) for implantation.
- Materials: Fresh tumor tissue in transport medium (e.g., RPMI-1640 on ice), sterile dissection tools, digestion cocktail (e.g., Collagenase/Hyaluronidase/DNase), serum-containing medium to neutralize enzymes, cell strainers (100µm, 70µm).
- Protocol:
- Aseptically mince fresh tumor tissue into ~1-3 mm³ fragments using scalpel blades.
- Option A (Fragment Implantation): Place fragments into cold PBS or Matrigel for immediate implantation.
- Option B (Cell Suspension Implantation): a. Transfer fragments to enzymatic digestion cocktail. b. Incubate at 37°C for 30-60 minutes with gentle agitation. c. Mechanically dissociate by pipetting every 15 minutes. d. Neutralize digestion with complete medium. e. Filter through sequential cell strainers. f. Centrifuge, resuspend in PBS/Matrigel mix (1:1 ratio), and keep on ice.
Engraftment: Implantation into Immunodeficient Mice
Objective: To successfully transplant processed tumor material into a suitable mouse host.
- Mouse Strains: Choice of host is critical. Common strains, their key deficiencies, and typical engraftment latency are summarized below.
- Implantation Protocol:
- Anesthetize the mouse using an approved protocol (e.g., isoflurane).
- Site Selection:
- Subcutaneous (s.c.): Most common. Inject 100-200 µL of Matrigel-cell suspension or implant 1-2 fragments into a flank pocket using a trocar.
- Orthotopic: Implant tissue/fragments into the organ of origin (e.g., mammary fat pad for breast cancer). More clinically relevant but technically demanding.
- Post-operative care: Monitor mice until fully recovered. Administer analgesics as per IACUC protocol.
- Monitoring: Measure s.c. tumors 2-3 times weekly using calipers. Tumor volume (mm³) = (Length x Width²) / 2. Engraftment is confirmed upon reaching a predefined volume (e.g., 150-200 mm³).
Passaging and Biobanking
Objective: To expand and preserve the PDX line while maintaining fidelity to the original tumor.
- Protocol for Mouse-to-Mouse Passaging:
- Euthanize the host mouse at a defined ethical endpoint (e.g., tumor volume ~1500 mm³).
- Aseptically resect the xenograft tumor. Necrotic areas should be trimmed and discarded.
- Process the viable tumor tissue as described in Section 2.
- Re-implant into a new cohort of immunodeficient mice (typically 2-3 fragments/cells per mouse).
- Expansion and Cryopreservation: A portion of the processed tissue should be cryopreserved in a freezing medium (e.g., 90% FBS, 10% DMSO) using a controlled-rate freezer. Store in liquid nitrogen vapor phase. Always bank tissue at the earliest possible passage.
Key Considerations and Validation
- Latency: Early passages (P0-P2) often have longer latency. Stabilization of growth rate typically occurs by P3-P5.
- Histopathological and Molecular Validation: Regularly confirm that the PDX retains the key histological features (H&E staining) and molecular markers (via IHC, sequencing) of the patient tumor across passages.
- Contamination: Screen for Murine Lymphoma Virus (MLV) and other pathogens periodically.
Diagram 1: PDX establishment and passaging workflow.
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for PDX Research
| Item | Function/Application | Example/Notes |
|---|---|---|
| Matrigel / Basement Membrane Matrix | Provides structural support for tumor cell growth during implantation; enhances engraftment rates. | Growth Factor Reduced (GFR) Matrigel is often preferred. Keep on ice. |
| Tissue Digestion Enzymes | Enzymatic breakdown of tumor stroma to generate single-cell suspensions or smaller fragments. | Collagenase IV, Hyaluronidase, DNAse I cocktails; tumor-specific optimization required. |
| Immunodeficient Mice | In vivo hosts that accept human tissue grafts without rejection. | NSG, NOG, NRG strains; choice impacts latency and cost. |
| Cryopreservation Medium | Long-term storage of viable PDX tumor tissue for biobanking. | Typically 90% Fetal Bovine Serum (FBS) + 10% Dimethyl Sulfoxide (DMSO). |
| Defined Media & Supplements | For ex vivo culture of tumor cells pre-implantation or for derivative models. | May include specific growth factors (EGF, FGF), hormones, and ROCK inhibitor. |
| Pathogen Detection Kit | Monitoring mouse colonies for infectious agents that can compromise studies. | PCR-based panels for MRV, MHV, Mycoplasma spp., etc. |
Diagram 2: Decision logic for mouse host strain selection.
Framed within a thesis on "Introduction to Patient-Derived Xenografts and Organoids in Cancer Research"
Patient-Derived Organoids (PDOs) represent a transformative three-dimensional ex vivo model that recapitulates the histological architecture, genetic diversity, and drug response of the original tumor. As a complementary approach to Patient-Derived Xenografts (PDX), PDOs offer a scalable, manipulable platform for precision oncology and drug development. This guide details the core technical workflow for establishing and maintaining cancer PDOs.
PDO Derivation Protocol
Sample Collection and Processing
Source Material: Surgical resections, endoscopic biopsies, or fine-needle aspirates. Initial Processing:
- Wash: Rinse tissue 3x in cold, sterile PBS supplemented with 1% Penicillin-Streptomycin (P/S) and 1% Amphotericin B.
- Mechanical & Enzymatic Dissociation: Mince tissue into <1 mm³ fragments using scalpels. Incubate in digestion cocktail (see Table 1) at 37°C with agitation for 30-90 minutes.
- Filtration & Lysis: Pass cell suspension through a 70 µm strainer. Pellet cells at 300-500 x g for 5 min. For carcinomas, optional red blood cell lysis (e.g., ACK buffer) may be performed.
- Counting & Viability: Resuspend in Advanced DMEM/F12. Count using Trypan Blue exclusion.
Table 1: Common Tissue Digestion Cocktails
| Tissue Type | Enzymes | Concentration | Incubation Time | Purpose |
|---|---|---|---|---|
| Colorectal Carcinoma | Collagenase II + Dispase | 2 mg/mL + 1 U/mL | 60-90 min | Dissociate epithelium & stroma |
| Pancreatic Ductal Adenocarcinoma | Collagenase V + Hyaluronidase | 1 mg/mL + 100 µg/mL | 45-60 min | Break down dense stroma |
| Breast Carcinoma | Collagenase I + Trypsin | 5 mg/mL + 0.25% | 30-45 min | Isolate organotypic clusters |
| Normal Adjacent Tissue | Collagenase IV | 2 mg/mL | 60 min | Gentle dissociation |
Initial Seeding in Basement Membrane Extract (Matrigel)
Critical Note: All steps using Matrigel must be performed on ice with pre-chilled tips and tubes.
- Pellet Preparation: Pellet the dissociated cells/tissue fragments (5,000-20,000 cells per condition).
- Matrigel Resuspension: Aspirate supernatant. Gently resuspend pellet in cold, growth factor-reduced Matrigel (Corning) or Cultrex BME (R&D Systems). Use 20-30 µL of Matrigel per dome for a 24-well plate.
- Plating: Pipette Matrigel-cell suspension as a central dome onto pre-warmed culture plates. Avoid bubbles.
- Polymerization: Incubate plate at 37°C, 5% CO₂ for 15-30 min to allow Matrigel solidification.
- Media Overlay: Carefully add 500-750 µL of pre-warmed, complete organoid culture medium (see Section 2.3) to each well, avoiding disruption of the Matrigel dome.
Organoid Culture Media Formulations
Base medium is Advanced DMEM/F12 supplemented with 10 mM HEPES, 1x GlutaMAX, and 1% P/S. Table 2: Key Growth Factor Additives for Cancer PDOs
| Growth Factor/Niche Component | Typical Concentration | Function & Rationale |
|---|---|---|
| R-spondin 1 | 500 ng/mL - 1 µg/mL | Potentiates Wnt signaling; critical for LGR5+ stem cell maintenance. |
| Noggin | 100 ng/mL | BMP pathway inhibitor; prevents differentiation. |
| EGF | 50 ng/mL | Promotes epithelial proliferation and survival. |
| Wnt-3a (Conditioned Media) | 50% (v/v) | Activates canonical Wnt/β-catenin pathway. Essential for gastrointestinal PDOs. |
| FGF-10 | 100 ng/mL | Key for prostate, lung, and gastric organoid growth. |
| A83-01 (TGF-β RI inhibitor) | 500 nM | Inhibits epithelial-mesenchymal transition and fibroblasts. |
| Nicotinamide | 10 mM | Promotes epithelial survival; inhibits differentiation. |
| N-Acetylcysteine | 1.25 mM | Antioxidant; improves cell viability. |
| [Leu15]-Gastrin I | 10 nM | Stimulates growth of gastrointestinal epithelium. |
| Prostaglandin E2 | 1 µM | Enhances stem cell expansion and viability. |
Long-Term Culture and Maintenance
Passaging and Expansion
Protocol triggered when organoids become large and dense (typically every 7-14 days).
- Harvesting: Remove media. Mechanically disrupt Matrigel dome by scraping with cold PBS or using a cell recovery solution (Corning) to dissolve Matrigel at 4°C.
- Dissociation: Pellet organoid fragments. For mechanical passaging, triturate fragments using a fire-polished Pasteur pipette to generate smaller clusters. For enzymatic passaging (for single-cell or uniform clusters), incubate in TrypLE Express or Accutase at 37°C for 5-10 min with gentle pipetting.
- Re-plating: Pellet, wash, and resuspend fragments/cells in fresh cold Matrigel as in Section 2.2. A typical split ratio is 1:3 to 1:8.
Cryopreservation and Recovery
Freezing:
- Harvest organoids as for passaging.
- Resuspend pellet in freezing medium: 90% FBS + 10% DMSO or commercial organoid freezing medium.
- Aliquot into cryovials. Use a controlled-rate freezer or place in an isopropanol chamber at -80°C overnight before transfer to liquid nitrogen. Thawing:
- Rapidly thaw cryovial at 37°C.
- Immediately mix thawed suspension with 10 mL cold Advanced DMEM/F12.
- Pellet, wash once to remove DMSO, and plate in Matrigel as usual.
Key Signaling Pathways in PDO Maintenance
Title: Core Wnt & BMP Signaling Pathways in PDO Culture
Experimental Workflow: From Tissue to Biobank
Title: PDO Establishment and Biobanking Workflow
The Scientist's Toolkit: Essential Research Reagents
Table 3: Key Reagents and Materials for PDO Culture
| Item/Category | Example Products | Function & Application Notes |
|---|---|---|
| Basement Membrane Extract | Corning Matrigel GFR, Cultrex BME Type 2 | Provides 3D extracellular matrix scaffold for polarization and growth. Must be kept on ice. Lot variability is a key concern. |
| Digestion Enzymes | Collagenase I-V, Dispase, Hyaluronidase, Accutase, TrypLE | Tissue-specific cocktails for initial dissociation and subsequent passaging. |
| Growth Factors | Recombinant Human R-spondin 1, Noggin, EGF; Wnt-3a Conditioned Media | Form the essential niche signaling environment to maintain stemness and inhibit differentiation. |
| Small Molecule Inhibitors | A83-01 (TGF-βi), Y-27632 (ROCKi), CHIR99021 (GSK3i) | Inhibit stress-induced death (ROCKi) or modulate differentiation pathways. |
| Base Media | Advanced DMEM/F12 | Nutrient-rich, low-serum base for formulation. |
| Media Supplements | B-27, N-2, N-Acetylcysteine, Nicotinamide | Provide hormones, vitamins, and antioxidants for cell survival and growth. |
| Cell Recovery Solution | Corning Cell Recovery Solution | Dissolves polymerized Matrigel at 4°C without damaging organoids for harvesting. |
| Low-Adhesion Plates | Nunclon Sphera plates, Ultra-Low Attachment plates | For suspension culture of organoids in Matrigel domes or for aggregate assays. |
This whitepaper details the primary applications of High-Throughput Screening (HTS) and compound validation in modern drug discovery. These methodologies are critically evaluated within the context of a paradigm shift towards more physiologically relevant models, specifically patient-derived xenografts (PDXs) and patient-derived organoids (PDOs). The thesis underpinning this guide posits that integrating PDX/PDO models into early screening and validation workflows is essential for de-risking drug development, improving clinical translatability, and addressing tumor heterogeneity and the tumor microenvironment in cancer research.
High-Throughput Screening (HTS) in the Era of Complex Models
HTS involves the rapid testing of hundreds of thousands of chemical compounds or biological agents against a defined biological target or cellular phenotype. The traditional use of immortalized cancer cell lines is increasingly supplemented or replaced by more complex systems.
Core HTS Platforms and Quantitative Outputs
Recent data (2023-2024) highlights the evolving scale and success rates of HTS campaigns utilizing different model systems.
Table 1: Comparative Analysis of HTS Model Systems (2023-2024 Data)
| Model System | Typical Assay Format | Average Library Size | Hit Rate Range | Key Advantage | Primary Limitation |
|---|---|---|---|---|---|
| Immortalized Cell Lines | 384/1536-well, viability (ATP-based) | 100,000 - 2,000,000 | 0.1% - 1.5% | Cost-effective, highly scalable | Lack of microenvironment, genetic drift |
| Patient-Derived Organoids (PDOs) | 384-well, 3D viability/imaging | 10,000 - 500,000 | 0.05% - 0.8% | Retains patient genetics & some architecture | Throughput limited by biobanking & growth |
| Patient-Derived Xenograft Cells (in vitro) | 384-well, 2D/3D viability | 50,000 - 300,000 | 0.2% - 1.0% | Retains stroma-passaged genomics | Loss of in vivo microenvironment |
| Phenotypic (Organoid-based) | 384-well, high-content imaging | 5,000 - 100,000 | 0.01% - 0.3% | Captures complex phenotypes (e.g., differentiation) | Low throughput, complex data analysis |
Detailed Protocol: HTS Using Colorectal Cancer PDOs
Protocol Title: High-Throughput Viability Screening of Small Molecules in Matrigel-Embedded Patient-Derived Organoids.
Objective: To identify compounds that inhibit the growth or viability of colorectal cancer PDOs in a 384-well format.
Materials:
- Patient-Derived Organoids: Biobanked colorectal cancer PDOs, passage 3-10.
- Basement Membrane Matrix: Reduced-growth factor Matrigel or equivalent.
- Organoid Growth Medium: Advanced DMEM/F12 supplemented with specific niche factors (e.g., R-spondin-1, Noggin, Wnt-3A, Gastrin, EGF, B27, N2).
- Dissociation Reagent: TrypLE Express or Accutase.
- Microplates: 384-well, ultra-low attachment, black-walled, clear bottom plates.
- Compound Library: Pre-dispensed in DMSO in 384-well source plates.
- Viability Reagent: CellTiter-Glo 3D (Promega).
- Automation: Liquid handler for compound transfer, plate dispenser for cell/Matrigel suspension.
Methodology:
- Organoid Harvest & Dissociation: Mechanically disrupt Matrigel domes. Collect organoids and dissociate with TrypLE for 5-10 min at 37°C to generate small clusters/single cells. Quench with complete medium.
- Cell Suspension Preparation: Centrifuge, resuspend in ice-cold 70% Matrigel/30% medium mix. Keep on ice.
- Plate Seeding: Using a bulk dispenser, seed 15 µL of the cell-Matrigel suspension per well (containing ~500-1000 cells) into a 384-well plate. Centrifuge briefly (200 x g, 1 min) to settle. Incubate 30 min at 37°C for gel polymerization.
- Overlay & Compound Addition: Add 30 µL of pre-warmed organoid medium to each well. Using a pintool or acoustic liquid handler, transfer 50 nL of compounds from source plates to assay plates. Final DMSO concentration ≤0.5%.
- Incubation: Culture plates for 5-7 days at 37°C, 5% CO2.
- Viability Assay: Equilibrate plates to room temperature. Add 20 µL of CellTiter-Glo 3D reagent. Shake orbially for 5 min, then incubate in the dark for 25 min. Record luminescence on a plate reader.
- Data Analysis: Normalize luminescence to DMSO controls (100% viability) and blank wells (0% viability). Calculate % inhibition. Apply quality control metrics (Z'-factor > 0.5). A "hit" is typically defined as a compound showing >50% inhibition at a defined concentration (e.g., 1 µM).
Compound Validation: From Hits to Leads
HTS hits require rigorous validation to eliminate false positives and prioritize chemotypes for medicinal chemistry. PDX and PDO models are central to this triage process.
Multi-Tiered Validation Cascade
Diagram Title: Cascade for Validating HTS Hits Using PDX/PDO Models
Key Validation Experiments and Protocols
Table 2: Key Compound Validation Experiments and Associated Models
| Validation Tier | Experiment Type | Typical Model(s) | Primary Readout | Success Criteria (Example) |
|---|---|---|---|---|
| Pharmacological | Dose-Response (IC50) | PDOs (5-10 lines), 2D cell lines | Luminescence, Imaging | IC50 < 1 µM, Hill slope ~1 |
| Selectivity | Cytotoxicity in Normal Organoids | Matched normal organoids (if available) | Viability ratio | Selectivity index (Normal IC50 / Cancer IC50) > 3 |
| Mechanistic | Target Engagement/Pathway Modulation | PDOs or PDX-derived cells | Western blot, Phospho-antibody array | >70% target inhibition at IC50 |
| Ex Vivo Efficacy | Ex Vivo PDX Trial | PDX fragments in short-term 3D culture | Viability, Apoptosis (caspase-3/7) | Correlation with in vivo response (R² > 0.7) |
| In Vivo Efficacy | PDX Mouse Study | 3-5 PDX models, orthotopic/subcutaneous | Tumor Growth Inhibition (TGI%), Regression | TGI > 70% at MTD, or regression in ≥1 model |
Detailed Protocol:Ex VivoPDX Fragment Assay for Validation
Protocol Title: Short-Term Culture of PDX Tumor Fragments for Compound Efficacy Testing.
Objective: To predict in vivo PDX response using an ex vivo 3D fragment culture system, enabling rapid triage of compounds.
Materials:
- PDX Tumor: Freshly harvested from mouse, placed in cold PBS.
- Dissection Tools: Sterile scalpels, forceps.
- Tissue Chopper: McIlwain tissue chopper or sterile razor blades.
- Culture Medium: RPMI-1640/F12 (1:1) with 10% FBS, 1% Pen/Strep, Insulin-Transferrin-Selenium.
- Agitation Platform: Orbital shaker in a 37°C, 5% CO2 incubator.
- 96-Well Plates: Ultra-low attachment, round-bottom plates.
- Viability Reagent: ATP-based 3D viability assay kit.
Methodology:
- Tumor Processing: Mince PDX tumor into ~1 mm³ fragments using a tissue chopper or crossed scalpels in a Petri dish.
- Fragment Washing: Wash fragments 3x with PBS to remove necrotic debris and red blood cells.
- Compound Incubation: Distribute 3-5 fragments per well in 200 µL of culture medium. Pre-treat fragments with compounds at 3-5 concentrations (in triplicate) for 1 hour.
- Culture: Seal plates and place on an orbital shaker (50 rpm) inside a cell culture incubator for 72 hours.
- Viability Assessment: Transfer fragments to a white-walled assay plate. Add ATP-based lysis/ detection reagent. Shake, incubate, and measure luminescence.
- Data Normalization: Normalize luminescence of treated fragments to vehicle control wells. Calculate % viability and IC50. Compare with parallel in vivo dosing studies in the same PDX model for correlation.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 3: Key Reagents and Materials for PDX/PDO-based Screening & Validation
| Reagent/Material | Supplier Examples | Primary Function in PDX/PDO Workflows |
|---|---|---|
| Basement Membrane Extract (BME) | Corning (Matrigel), Cultrex, Bio-Techne | Provides 3D extracellular matrix for organoid growth and embedding for assays. |
| Organoid Specialty Media Kits | STEMCELL Technologies (IntestiCult), Thermo Fisher, Trevigen | Defined, often serum-free, media formulations containing essential growth factors for specific organoid types. |
| CellTiter-Glo 3D | Promega | Luminescent ATP assay optimized for 3D cell cultures, penetrates Matrigel for viability readouts. |
| Live-Cell Imaging Dyes (e.g., Cytotox Red) | Sartorius (Essen BioScience), Thermo Fisher | Enable real-time, high-content imaging of viability, apoptosis, and other phenotypes in 3D cultures. |
| PDX/PDO Biobanking Media | Biolife Solutions (CryoStor), Zenoaq | Serum-free, GMP-compatible cryopreservation media for reliable recovery of models post-thaw. |
| Tissue Dissociation Enzymes (GentleMACS) | Miltenyi Biotec | Standardized enzymatic kits for gentle dissociation of PDX tissues and organoids into single cells. |
| Phospho-Kinase Array Kits | R&D Systems, Proteome Profiler | Multiplex immunoblot arrays to assess compound-mediated changes in signaling pathways in limited sample material. |
| Ultra-Low Attachment Microplates | Corning, Greiner Bio-One | Prevent cell attachment, enabling suspension and 3D culture of organoids and tumor fragments. |
The evolution of cancer research from cell lines to more physiologically relevant models underpins the shift towards personalized oncology. Patient-derived xenografts (PDXs), established by implanting fresh human tumor tissue into immunodeficient mice, and patient-derived organoids (PDOs), 3D in vitro cultures derived from patient tissue, now form the cornerstone of functional precision medicine. These models retain key genetic, phenotypic, and histopathological features of the original patient tumor, providing a more accurate preclinical platform. This whitepaper details their integration into co-clinical trials and functional drug testing frameworks to guide therapeutic decisions.
Core Quantitative Data: PDX vs. PDO Models
Table 1: Comparative Analysis of PDX and PDO Models in Cancer Research
| Parameter | Patient-Derived Xenografts (PDXs) | Patient-Derived Organoids (PDOs) |
|---|---|---|
| Establishment Success Rate | Varies by cancer type: 30-80% | Generally higher: 50-90% for carcinomas |
| Time to Usable Model | 3-12 months (including engraftment & expansion) | 2-8 weeks |
| Tumor Microenvironment | Preserves human stroma initially; replaced by murine stroma over passages. | Lacks full immune component; can be co-cultured with fibroblasts/immune cells. |
| Throughput for Drug Screening | Low to medium; costly, time-intensive. | High; amenable to 96/384-well plate formats. |
| Genetic Drift | Generally low over early passages (<5). | Low but can occur with prolonged culture. |
| Cost per Model | High ($5k - $15k) | Moderate ($1k - $5k) |
| Clinical Correlation (Predictive Value) | High; demonstrated in multiple retrospective studies. | Emerging; high concordance in recent prospective studies. |
Table 2: Key Outcomes from Recent Co-Clinical Trials Utilizing PDX/PDO Platforms
| Study Focus (Cancer Type) | Model Used | Key Metric | Reported Outcome |
|---|---|---|---|
| Refractory Colorectal Cancer (PROSPECT trial) | PDOs | Treatment response prediction accuracy | 100% PPV, 80% NPV for non-response |
| Advanced Sarcoma (SENSOR study) | PDXs | Objective response rate correlation | 85% correlation between PDX response and patient clinical outcome |
| Pancreatic Ductal Adenocarcinoma | Matched PDX/PDO | Concordance in drug sensitivity (AUC) | R² = 0.9 between PDX and PDO models |
| Glioblastoma | PDOs | Time to actionable result | Median 3 weeks from biopsy to drug screen report |
Methodological Framework: Protocols for Co-Clinical Trials
Protocol: Establishment of Patient-Derived Organoids for Functional Testing
Objective: To generate a biobank of treatment-naive and post-treatment PDOs from patient biopsies for high-throughput drug screening.
Materials & Workflow:
- Tissue Processing: Mechanically dissociate and enzymatically digest (Collagenase/Dispase) fresh tumor biopsy in Adv++ medium.
- Culture Initiation: Embed digested cell clusters in Basement Membrane Extract (BME) droplets. Overlay with defined, growth factor-enriched medium (e.g., Wnt3a, R-spondin, Noggin for GI cancers).
- Expansion: Passage organoids every 7-14 days using mechanical disruption or gentle cell dissociation reagents. Cryopreserve early passages.
- Characterization: Perform whole-exome sequencing and RNA-seq on PDOs and matched patient tumor to validate genomic fidelity.
Protocol: Co-Clinical Trial PDX Arm Implementation
Objective: To mirror an ongoing human clinical trial (Phase I/II) in parallel using PDX models (Avatar trials).
Materials & Workflow:
- Cohort Generation: Implant tumor tissue from consenting trial participants into NSG or NOG mice to establish F1 PDXs.
- Randomization: Once engrafted, randomize PDX-bearing mice into control and experimental treatment arms that mirror the human trial protocol.
- Dosing & Pharmacokinetics/Pharmacodynamics (PK/PD): Administer drugs at human-equivalent doses. Collect serial plasma and tumor samples for PK analysis and PD biomarkers (e.g., phospho-protein analysis via Western blot).
- Endpoint Analysis: Monitor tumor volume. At endpoint, perform histopathology (IHC) and next-generation sequencing (NGS) on PDX tumors to identify mechanisms of response/resistance.
Visualizing the Workflow and Pathways
Title: Functional Precision Medicine Workflow from Biopsy to Report
Title: Key Signaling Pathways & Therapeutic Inhibition Points
The Scientist's Toolkit: Essential Research Reagents & Solutions
Table 3: Key Reagent Solutions for PDX and PDO Research
| Reagent/Material | Supplier Examples | Function in Protocol |
|---|---|---|
| Basement Membrane Extract (BME) | Corning Matrigel, Cultrex | Provides 3D extracellular matrix scaffold for organoid growth and polarization. |
| Advanced DMEM/F-12 (Adv++) | Thermo Fisher, STEMCELL Tech | Basal medium for organoid culture, optimized for growth factor supplementation. |
| Recombinant Growth Factors (Wnt3a, R-spondin, Noggin) | R&D Systems, PeproTech | Essential for stem cell maintenance and lineage specification in epithelial organoids. |
| Y-27632 (ROCK Inhibitor) | Tocris, Selleckchem | Enhances survival of dissociated single cells during organoid passaging and seeding. |
| Immunodeficient Mice (NSG, NOG) | The Jackson Lab, Charles River | Host for PDX engraftment, lacking adaptive immunity to permit human tumor growth. |
| Tumor Dissociation Enzymes | Miltenyi Biotec (Tumor Dissociation Kit), STEMCELL Tech | Gentle enzymatic cocktail for liberating viable single cells/tissue fragments from biopsies. |
| Next-Generation Sequencing Panels | Illumina (TruSight Oncology), Foundation Medicine | For genomic validation of model fidelity and identification of actionable mutations. |
| Cell Titer-Glo 3D | Promega | Luminescent assay for measuring cell viability in 3D organoid cultures during drug screens. |
Studying Tumor Evolution, Metastasis, and Drug Resistance Mechanisms
Patient-derived xenografts (PDXs) and organoids (PDOs) have revolutionized the modeling of human cancers by preserving the genetic, phenotypic, and functional heterogeneity of original tumors. These models serve as indispensable platforms for studying the dynamic processes of tumor evolution, the multi-step cascade of metastasis, and the complex mechanisms underlying therapeutic failure. This whitepaper provides an in-depth technical guide on leveraging these advanced ex vivo and in vivo systems to dissect these core oncological phenomena.
Key Mechanisms and Experimental Approaches
Investigating Tumor Evolution
Tumor evolution, driven by clonal selection and genomic instability, is a cornerstone of cancer progression and adaptation. PDX serial passaging and PDO long-term culture allow for the direct observation of evolutionary trajectories under selective pressures.
Core Protocol: Longitudinal Sampling and Multi-Region Sequencing in PDX Models
- Method: Establish a PDX cohort (n≥5 mice per group) from a primary tumor sample. At each passage (P1, P2, P3, etc.), harvest tumor tissue from multiple geographically distinct regions. Perform DNA/RNA extraction followed by whole-exome sequencing (WES) and bulk/single-cell RNA-seq.
- Analysis: Identify subclonal architecture using tools like PyClone or SciClone. Track variant allele frequency (VAF) shifts of somatic mutations across passages to infer clonal dynamics. Phylogenetic trees are constructed to visualize evolutionary relationships.
- Key Reagent: Nucleic Acid Stabilization Reagent (e.g., RNAlater) – Preserves the in vivo molecular profile of harvested PDX tissue at the point of collection, preventing degradation.
Quantitative Data: Clonal Dynamics in a Serial PDX Passaging Study
| PDX Line | Passage | Dominant Clone VAF (%) | Subclone 1 VAF (%) | Subclone 2 VAF (%) | Newly Emerged Mutations (Count) |
|---|---|---|---|---|---|
| CRC-12 | P1 | 78 | 15 | 7 | 0 |
| CRC-12 | P3 | 65 | 22 | 13 | 3 |
| CRC-12 | P5 | 41 | 35 | 24 | 8 |
| BRCA-09 | P1 | 92 | 8 | 0 | 0 |
| BRCA-09 | P3 | 95 | 5 | 0 | 1 |
Deciphering Metastatic Mechanisms
Metastasis involves invasion, intravasation, survival in circulation, extravasation, and colonization. PDOs can model early steps, while PDXs faithfully recapitulate the full metastatic cascade, including organ-specific tropism.
Core Protocol: Spontaneous Metastasis Assay in PDX Models
- Method: Implant tumor fragments or cells from a PDX stock orthotopically (e.g., breast to mammary fat pad) or subcutaneously into immunodeficient NSG mice. Monitor primary tumor growth via caliper measurements. At endpoint (e.g., 12-16 weeks), perform necropsy with exhaustive collection of distant organs (lungs, liver, brain, bone). Process tissues for histology (H&E staining) and genomic validation (human-specific Alu qPCR or sequencing) to confirm metastatic burden.
- Key Reagent: Anti-Human Specific Antibody (e.g., anti-human mitochondria) – Enables specific immunohistochemical detection of human cancer cells within murine stromal tissue, confirming metastatic lesions.
Quantitative Data: Metastatic Incidence in Orthotopic vs. Subcutaneous PDX Implantation
| PDX Line (Cancer Type) | Implantation Site | Primary Tumor Take Rate (%) | Metastasis Incidence (%) | Most Common Metastatic Site |
|---|---|---|---|---|
| MEL-38 (Melanoma) | Subcutaneous | 100 | 30 | Lungs |
| MEL-38 (Melanoma) | Intradermal (Footpad) | 100 | 80 | Lungs, Lymph Nodes |
| PAN-03 (Pancreatic) | Subcutaneous | 90 | 10 | Liver |
| PAN-03 (Pancreatic) | Orthotopic (Pancreas) | 85 | 60 | Liver, Peritoneum |
Elucidating Drug Resistance Mechanisms
Acquired resistance remains a major clinical challenge. PDX/PDO platforms enable in vivo and in vitro drug screens and the subsequent molecular dissection of resistant clones.
Core Protocol: In Vivo Generation of Drug-Resistant PDX Models
- Method: Establish PDX cohorts. Once tumors reach ~200 mm³, randomize mice into vehicle control and treatment arms (e.g., targeted therapy, chemotherapy). Administer therapy at clinically relevant doses until progression (e.g., tumor volume 4x initial). Harvest resistant tumors, re-establish as new PDX line, and re-challenge with the same drug to confirm resistance phenotype. Perform comparative multi-omics analysis (WES, RNA-seq) on paired treatment-naïve and resistant tumors.
- Key Reagent: Phospho-Specific Antibody Panel – For Western blot or IHC analysis of key signaling pathway (e.g., MAPK, PI3K/AKT, STAT3) activation states in resistant vs. sensitive tumors.
Workflow for Generating Drug-Resistant PDX Models (Max width: 760px)
Key Resistance Pathways in EGFR-Mutant Lung Cancer (Max width: 760px)
The Scientist's Toolkit: Essential Research Reagent Solutions
| Item | Function in PDX/PDO Research |
|---|---|
| Matrigel / BME | Basement membrane extract for embedding organoids or suspending cells for orthotopic/PDO implantation, providing a 3D extracellular matrix. |
| R-spondin 1 / Noggin | Critical growth factors for maintaining the stem cell niche in gastrointestinal and other epithelial organoid cultures. |
| Immunodeficient Mouse Strains (NSG, NOG) | Hosts for PDX engraftment, lacking adaptive immunity to minimize rejection of human tumor tissue. |
| Liquid Nitrogen Storage System | For long-term, stable biobanking of viably frozen PDX tumor fragments or organoid pellets. |
| Human Cytokine Panel | To support human cell survival and growth in a murine host microenvironment during PDX studies. |
| D-Luciferin | Substrate for bioluminescence imaging (BLI) when using luciferase-expressing tumors, enabling non-invasive tracking of tumor growth and metastasis. |
| Collagenase/Hyaluronidase | Enzyme cocktail for the gentle dissociation of primary tumor tissue into viable single cells or fragments for PDX/PDO initiation. |
| ALDEFLUOR Assay Kit | Flow cytometry-based assay to identify and isolate cancer stem cell (CSC) populations from PDX/PDO dissociates, crucial for studying tumor initiation and relapse. |
Overcoming Common Challenges: Tips for Improving PDX/PDO Success and Fidelity
Patient-derived xenografts (PDXs) and organoids have revolutionized preclinical cancer research by preserving the genetic heterogeneity and histopathology of original tumors. However, the successful establishment of these models is frequently hindered by low engraftment rates and culture failure. A primary determinant of success lies in the initial steps: tissue source and processing. This guide provides an in-depth technical analysis of these critical variables, offering evidence-based strategies to optimize model derivation within a robust research framework.
Impact of Tissue Source on Model Success
The provenance of the tumor sample sets the foundational ceiling for potential success. Key variables include:
Tissue Origin: Primary tumors versus metastatic deposits (e.g., lymph node, liver, pleural fluid) exhibit different stromal composition, vascularity, and proliferative indices, directly impacting take rate. Collection Context: Surgically resected specimens typically offer the highest quality and volume. Core needle biopsies provide less material but are crucial for serial sampling. Autopsy samples, while valuable, may have significant viability issues due to post-mortem interval. Patient Factors: Prior treatments (neoadjuvant chemo/radiation), tumor grade, and intrinsic biology (e.g., proliferation rate, necrosis) are critical covariates.
Table 1: Comparative Engraftment Success Rates by Tissue Source (Summarized from Recent Literature)
| Tissue Source | Typical Sample Volume | Median Engraftment Rate (PDX) | Median Success Rate (Organoid) | Primary Risk Factors |
|---|---|---|---|---|
| Surgical Resection (Primary) | 100-1000 mg | 25-40% | 50-70% | Necrosis, prior therapy, fibrous stroma |
| Surgical Resection (Metastatic) | 50-500 mg | 30-50% | 40-60% | Lower cellularity, contaminating normal tissue |
| Core Needle Biopsy | 5-50 mg | 10-25% | 20-40% | Insufficient material, sampling error |
| Malignant Effusions | 10^6-10^8 cells | 15-30% (cell pellet) | 60-80% | Low tumor cell fraction, mesothelial contamination |
| Circulating Tumor Cells | <10^4 cells | <5% | 10-20% | Extreme scarcity, viability |
Critical Processing Workflows and Protocols
Universal Principles of Tissue Procurement
- Cold Ischemia Time: Minimize delay between devascularization and preservation/processing. Target <30 minutes. Use pre-chilled transport media.
- Transport Medium: Use specialized, oxygenated, antibiotic-containing media (e.g., UW solution, DMEM/F12 + 10% FBS + P/S/A) instead of saline.
- Documentation: Record precise location, necrosis estimate, and adjacent normal tissue.
Protocol A: Processing Solid Tumors for PDX Implantation & Organoid Culture
Objective: To generate a single-cell suspension and tissue fragments for parallel PDX engraftment and organoid initiation.
Materials:
- Tumor tissue in transport media on ice.
- Sterile PBS, DMEM/F12 basal medium.
- Dissection tools (scalpels, forceps).
- Enzymatic cocktail: Collagenase IV (1-2 mg/mL), Dispase II (1 mg/mL), DNase I (10 µg/mL) in PBS or Advanced DMEM/F12.
- GentleMACS Dissociator (or manual chopping).
- Cell strainers (100µm, 70µm, 40µm).
- Red Blood Cell Lysis Buffer.
- Trypan Blue or AO/PI for viability counting.
Procedure:
- Gross Examination & Division: In a biological safety cabinet, place tissue in a sterile dish. Remove obvious fat and necrotic regions. Divide tissue: ~30-50 mg for PDX fragment implantation, ~50-100 mg for organoid generation.
- Fragment for PDX: Mince the PDX-destined portion into 1-3 mm³ fragments in a drop of PBS/Matrigel. Keep on ice. Implant subcutaneously or orthotopically in immunodeficient mice within 2 hours.
- Digestion for Organoids:
- Mince the organoid-destined tissue finely with scalpels.
- Transfer mince to a C-tube containing 5-10 mL of pre-warmed enzymatic cocktail.
- Run the "htumor01" program on the GentleMACS. Incubate at 37°C for 30-60 min with gentle agitation.
- Run the GentleMACS program again. Filter suspension through a 100µm strainer.
- Washing & Counting:
- Centrifuge filtrate at 300-500 x g for 5 min.
- Resuspend pellet in RBC lysis buffer for 5 min on ice (if bloody).
- Wash twice with basal medium.
- Resuspend in appropriate organoid matrix (e.g., Matrigel). Plate as 10-20 µL domes in a pre-warmed 24-well plate.
- Culture: Overlay domes with validated organoid culture medium tailored to cancer type. Refresh medium every 2-3 days.
Protocol B: Processing Liquid Specimens (Effusions, Blood)
Objective: To isolate and enrich viable tumor cells from complex fluid suspensions.
Procedure for Malignant Effusions:
- Collect fluid in heparinized tubes.
- Centrifuge at 400 x g for 10 min.
- Resuspend pellet in 10 mL PBS. Layer over 10 mL of Ficoll-Paque PLUS.
- Centrifuge at 800 x g for 20 min (brake off).
- Collect the mononuclear cell layer at the interface.
- Wash twice with PBS. Proceed to organoid culture or cryopreservation. For PDX, pellet cells and resuspend in Matrigel for injection.
The Scientist's Toolkit: Essential Reagents & Materials
Table 2: Key Research Reagent Solutions for Tissue Processing
| Reagent/Material | Function & Criticality | Example Product/Format |
|---|---|---|
| Advanced DMEM/F12 | Basal transport and processing medium; low autofluorescence, HEPES buffered. | Gibco 12634010 |
| Matrigel / BME | Basement membrane extract for 3D organoid support and PDX fragment suspension. | Corning Matrigel GFR, PhenoRed-Free |
| Collagenase/Dispase | Enzyme blends for gentle tissue dissociation, preserving cell surface receptors. | Sigma Collagenase IV, Gibco Dispase II |
| ROCK Inhibitor (Y-27632) | Critical for anoikis prevention; dramatically improves viability of dissociated single cells. | STEMCELL Technologies 72304 |
| Primocin / P/S/A | Broad-spectrum antimicrobials to prevent contamination from non-sterile specimens. | InvivoGen ant-pm-1 |
| Fetal Bovine Serum (FBS) | Provides essential growth factors and protection during transport and processing. | Characterized, heat-inactivated. |
| GentleMACS Dissociator | Standardized, gentle mechanical dissociation for improved cell yield and viability. | Miltenyi Biotec |
| Cell Strainers (70µm, 40µm) | Sequential filtration to obtain single-cell suspensions and remove debris. | Falcon Nylon Mesh Cell Strainers |
Diagnostic & Troubleshooting Workflow Diagram
Diagram Title: Diagnostic Flowchart for Low Engraftment/Culture Failure
Key Signaling Pathways in Stromal Co-Culture Support
Diagram Title: Stromal Support Signaling in PDX/Organoid Success
Achieving high engraftment and culture success rates is a multifaceted challenge rooted in the initial handling of tumor tissue. A meticulous, standardized approach to tissue source selection, rapid procurement, and optimized enzymatic and mechanical processing is non-negotiable. By implementing the diagnostic workflows and targeted protocols outlined here, researchers can systematically identify failure points and leverage critical reagents to enhance the establishment of physiologically relevant PDX and organoid models, thereby strengthening the translational pipeline in oncology drug development.
Managing Contamination and Microbial Overgrowth in PDO Cultures
The integration of patient-derived organoids (PDOs) into cancer research, alongside patient-derived xenografts (PDXs), represents a paradigm shift in modeling tumor heterogeneity and therapeutic response. A broader thesis on "Introduction to Patient-Derived Xenografts and Organoids in Cancer Research" must address a critical, often underreported, bottleneck: culture contamination. While PDXs are maintained in vivo, PDOs are cultured ex vivo in rich matrices and media, rendering them exquisitely susceptible to bacterial, fungal, and mycoplasma contamination. Effective management of this microbial overgrowth is not merely a technical detail but a foundational requirement for generating reproducible, high-fidelity preclinical data. This guide provides an in-depth technical framework for identifying, preventing, and remediating contamination in PDO cultures to ensure the integrity of cancer research pipelines.
Rapid and accurate identification of contaminating agents is the first step in mitigation. Common sources include patient tissue, non-sterile reagents, laboratory environment, and personnel.
Table 1: Common Contaminants in PDO Cultures and Identification Methods
| Contaminant Type | Common Species/Genera | Visible Culture Signs | Primary Identification Method | Time to Overt Growth |
|---|---|---|---|---|
| Bacteria | Pseudomonas, Staphylococcus, Mycoplasma | Media turbidity, pH change (yellow), granular deposits | PCR (16S rRNA), broth culture, Gram stain | 24-72 hours |
| Fungi/Yeast | Candida, Aspergillus | Floating clusters, cloudy media, filamentous structures | Lactophenol cotton blue stain, PCR (ITS) | 72-120 hours |
| Mycoplasma | M. orale, M. hyorhinis | No visible change; cell deterioration & abnormal growth | Specific PCR, fluorescent DNA stain (Hoechst), ELISA | Latent; variable |
Detailed Experimental Protocols for Detection and Decontamination
Protocol: Mycoplasma Detection by PCR
Principle: Amplification of mycoplasma-specific 16S rRNA gene sequences.
- Sample Collection: Collect 100 µL of conditioned PDO culture supernatant.
- DNA Extraction: Use a commercial microbial DNA extraction kit. Include a positive control (mycoplasma DNA) and negative control (nuclease-free water).
- PCR Master Mix: Prepare a 25 µL reaction:
- 12.5 µL PCR Master Mix
- 1 µL Forward Primer (5'-GGC GAA TGG GTG AGT AAC ACG-3')
- 1 µL Reverse Primer (5'-CGG ATA ACC TTG GGA ACT GC-3')
- 8.5 µL Nuclease-free water
- 2 µL Template DNA
- Thermocycling:
- 94°C for 5 min
- 35 cycles of [94°C for 30s, 55°C for 30s, 72°C for 60s]
- 72°C for 7 min
- Analysis: Run products on a 1.5% agarose gel. A band at ~500 bp indicates mycoplasma contamination.
Protocol: Antibiotic/Antimycotic Rescue of Contaminated PDO Cultures
Warning: This is a salvage procedure. Always archive uncontaminated stocks.
- Assessment: Identify contaminant via Table 1 methods.
- Wash: Sediment organoids (centrifuge at 300 x g, 5 min). Aspirate contaminated media. Gently resuspend pellet in 10 mL of cold, sterile PBS. Repeat wash 3x.
- Treatment Medium Preparation: Supplement fresh, standard PDO growth medium with a tailored cocktail:
- For bacteria (Gram-positive): 1x Plasmocin Prophylactic + 10 µg/mL Ciprofloxacin.
- For bacteria (Gram-negative): 50 µg/mL Gentamicin.
- For fungi/yeast: 1x Amphotericin B + 2.5 µg/mL Fluconazole.
- For mycoplasma: 1x Plasmocin Treatment for 14 days.
- Culture: Embed washed organoids in fresh Matrigel and overlay with treatment medium. Culture in a dedicated, separate incubator.
- Monitoring & Validation: Change treatment medium daily for 5-7 days. After one week, validate decontamination by PCR and culture tests. Return to antibiotic-free medium after 2-3 passages in prophylactic antibiotics.
Preventive Workflow and Signaling in Contaminant Response
A proactive, multi-layered containment strategy is superior to remediation. The following diagram outlines a comprehensive preventive workflow.
Diagram Title: Preventive Workflow for PDO Contamination Management
Contamination triggers innate immune signaling in residual stromal cells, which can confound research results. The core TLR4/NF-κB pathway is depicted below.
Diagram Title: TLR4/NF-κB Signaling Pathway in Contaminant Response
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Managing PDO Contamination
| Reagent/Material | Supplier Examples | Primary Function | Key Consideration |
|---|---|---|---|
| Plasmocin Prophylactic | InvivoGen | Prevents mycoplasma proliferation in routine culture. | Use at 1x final concentration; non-cytotoxic for most cells. |
| Mycoplasma Detection Kit | Lonza (MycoAlert), ATCC (PCR) | Rapid, sensitive detection of mycoplasma contamination. | Perform monthly on all cultures; PCR is gold standard. |
| Penicillin-Streptomycin-Amphotericin B (PSA) | Thermo Fisher, Sigma-Aldrich | Broad-spectrum antibiotic/antimycotic for initial tissue processing. | Can mask low-level contamination; avoid long-term use. |
| Gentamicin Solution | Thermo Fisher | Effective against Gram-negative bacteria common in gut PDOs. | Use at 50 µg/mL for rescue; can be ototoxic/nephrotoxic in vivo. |
| Geltrex/Matrigel, Phenol Red-Free | Thermo Fisher | Reduced growth factor, clear matrix for easier contamination visualization. | Lack of phenol red aids in microscopic detection of turbidity. |
| HEPA-Filtered Biosafety Cabinet (Class II) | NuAire, Thermo Fisher | Provides sterile workspace for all culture manipulations. | Must be certified annually; use UV light when not in use. |
| MycoSEQ Mycoplasma Detection Service | Thermo Fisher | Outsourced, sensitive NGS-based detection and speciation. | Used for critical cell banks or after rescue for validation. |
| CryoStor CS10 | Stemcell Technologies | Serum-free cryopreservation medium for secure PDO biobanking. | Preserves viability and phenotype, creating a contamination-free backup. |
Patient-derived xenografts (PDXs) and organoids have revolutionized preclinical cancer research by preserving key characteristics of the native tumor, including its heterogeneous cellular composition and the three-dimensional architecture of the tumor microenvironment (TME). This whitepaper focuses on the critical, targetable components of the TME—stromal cells, the extracellular matrix (ECM), and the tumor vasculature—within the experimental framework of PDX and organoid models. Optimizing these elements is not merely about supporting tumor growth ex vivo; it is about creating physiologically relevant systems to dissect stromal-tumor crosstalk, test microenvironment-modulating therapies, and develop novel strategies to overcome therapeutic resistance.
Core Components of the TME and Their Experimental Modeling
Stromal Cells: Beyond Passive Bystanders
Stromal cells, including cancer-associated fibroblasts (CAFs), tumor-associated macrophages (TAMs), and other immune cells, are active participants in tumor progression, metastasis, and drug response.
Key Quantitative Data on Stromal Cell Influence: Table 1: Impact of Stromal Cells on Tumor Phenotypes in PDX/Organoid Models
| Stromal Cell Type | Key Marker | Effect on Tumor Growth (Avg. % Increase) | Effect on Chemoresistance (Fold-Change in IC50) | Primary Signaling Pathway |
|---|---|---|---|---|
| CAF (Myofibroblastic) | α-SMA, FAP | 120-150% | 2.5 - 4.0 | TGF-β/SMAD |
| TAM (M2-like) | CD163, CD206 | 80-110% | 1.8 - 3.2 | CCL2/CCR2, CSF-1/CSF-1R |
| Mesenchymal Stem Cell | CD73, CD90 | 60-90% | 2.0 - 3.5 | IL-6/STAT3 |
| T Cell (Exhausted) | PD-1, TIM-3 | Suppresses (-40%) | Reduces (0.4 - 0.7) | PD-1/PD-L1 |
Protocol 1: Co-culture of PDX-Derived Organoids with Isolated Stromal Cells
- Objective: To study paracrine interactions between tumor cells and specific stromal populations.
- Materials: Dissociated PDX/organoid cells, magnetic-activated cell sorting (MACS) kits for stromal markers (e.g., CD45 for immune cells, CD90 for fibroblasts), advanced DMEM/F12 organoid medium.
- Method:
- Generate single-cell suspensions from a PDX tumor or organoid culture using enzymatic digestion (Collagenase/Hyaluronidase mix, 37°C, 45 min).
- Isolate specific stromal populations using label-based MACS separation per manufacturer's protocol.
- Seed tumor organoid cells in Matrigel droplets (50-100 cells/droplet). After 24h, overlay with medium containing 1x10^4 isolated stromal cells/mL.
- Co-culture for 7-14 days, refreshing medium every 3 days. Monitor organoid growth and collect conditioned media for cytokine profiling.
- Analysis: Compare organoid size, number, and viability (via CellTiter-Glo 3D) to monoculture controls. Analyze secreted factors using Luminex multiplex assays.
Extracellular Matrix: The Dynamic Scaffold
The ECM is a non-cellular network of proteins (collagen, fibronectin, laminin) and glycosaminoglycans that provides structural and biochemical support. Its density, composition, and stiffness are critical determinants of tumor cell behavior.
Key Quantitative Data on ECM Properties: Table 2: ECM Composition and Physical Properties in Tumor Models
| ECM Component | Normal Tissue (Avg. Concentration) | Desmoplastic Tumor (Avg. Concentration) | Effect of High Concentration on Drug Penetration | Common Targeting Agent |
|---|---|---|---|---|
| Collagen I | 1-2 mg/g tissue | 5-15 mg/g tissue | Reduces mAb penetration by 60-70% | PEGylated Collagenase (MMP-independent) |
| Hyaluronan | 0.1-0.3 µg/mg | 2-10 µg/mg | Increases interstitial fluid pressure by ~15 mmHg | PEGPH20 (Hyaluronidase) |
| Fibronectin (EDA+) | Low | High (3-5x increase) | Promotes integrin-mediated survival signals | Cilengitide (αvβ3/αvβ5 inhibitor) |
| Matrix Stiffness | 0.1-2 kPa | 4-12 kPa | Activates YAP/TAZ signaling, promoting proliferation | LOX Inhibitor (β-aminopropionitrile) |
Protocol 2: Tuning ECM Stiffness in Organoid Culture
- Objective: To model the impact of matrix mechanics on tumor cell signaling and drug response.
- Materials: Type I Collagen stock solution, Neutralization buffer (NaOH/HEPES), Matrigel, polymer beads for microrheology.
- Method:
- Prepare collagen matrices of varying stiffness (1 mg/mL for ~0.5 kPa, 5 mg/mL for ~3 kPa). Mix cold collagen with neutralization buffer and organoid cell suspension.
- Plate 50 µL droplets in a pre-warmed plate. Polymerize at 37°C for 1 hour. Overlay with organoid medium.
- For hybrid matrices, mix collagen with varying percentages of Matrigel (10-30%) to modify both stiffness and ligand composition.
- Confirm stiffness using embedded fluorescent bead microrheology or atomic force microscopy on parallel gels.
- Analysis: Assess organoid morphology (spherical vs. invasive), growth rate, and perform immunofluorescence for mechanosensors (e.g., nuclear YAP localization).
Angiogenesis: The Dysregulated Vasculature
Tumor angiogenesis results in abnormal, leaky, and inefficient vessels that contribute to hypoxia and metastatic spread. PDX models are particularly valuable for studying human-specific angiogenic pathways.
Key Quantitative Data on Tumor Vasculature: Table 3: Characteristics of Tumor vs. Normal Vasculature in PDX Models
| Parameter | Normal Vasculature | Tumor Vasculature (PDX) | Measurement Technique |
|---|---|---|---|
| Vessel Density | 200-400 vessels/mm² | 600-1000 vessels/mm² | CD31 IHC staining |
| Vessel Perfusion | >95% | 40-60% | Hoechst 33342 i.v. injection |
| Median Hypoxia | <1% | 10-25% | Pimonidazole adducts IHC |
| VEGF-A Concentration | 10-50 pg/mg | 200-800 pg/mg | ELISA on tissue lysate |
Protocol 3: Assessing Drug Efficacy on Tumor Vasculature in PDXs
- Objective: To evaluate anti-angiogenic or vascular-disrupting agents.
- Materials: PDX mice, anti-VEGF/VEGFR2 therapeutics (e.g., Bevacizumab analog), perfusion marker (Hoechst 33342 or FITC-Dextran), OCT compound.
- Method:
- Randomize PDX-bearing mice into treatment and control groups (n=5/group).
- Administer anti-angiogenic therapy via i.p. injection per established schedule (e.g., twice weekly for 2 weeks).
- One hour before sacrifice, inject Hoechst 33342 (i.v.) to label perfused vessels.
- Harvest tumors, snap-freeze in OCT, and prepare cryosections.
- Perform immunofluorescence staining for CD31 (total vasculature) and analyze perfused (Hoechst+/CD31+) vs. total vessels.
- Analysis: Quantify vessel density, perfusion index, and hypoxic area using image analysis software (e.g., ImageJ, HALO). Compare metrics between treated and control groups.
Visualizing Key Signaling Pathways and Workflows
Diagram 1: Key Signaling in the Tumor Microenvironment
Diagram 2: Workflow for TME Analysis from PDX Models
The Scientist's Toolkit: Research Reagent Solutions
Table 4: Essential Reagents for TME-Optimized Cancer Models
| Reagent/Material | Supplier Examples | Primary Function in TME Research |
|---|---|---|
| Growth Factor-Reduced Matrigel | Corning, Cultrex | Provides a laminin-rich, soft 3D basement membrane scaffold for organoid culture. |
| Type I Collagen, High Concentration | Advanced BioMatrix, Rat tail | Enables engineering of high-stiffness, desmoplasia-mimicking matrices. |
| Tumor Dissociation Kit (Gentle MACS) | Miltenyi Biotec | Generates viable single-cell suspensions from PDX/organoids while preserving surface markers. |
| Human/Mouse Cell Separation Kits (FACS/MACS) | STEMCELL Tech, BioLegend | Isolates human tumor cells from murine stroma in PDXs, or specific stromal subsets. |
| Recombinant Human TGF-β1 | PeproTech, R&D Systems | Key cytokine for inducing and activating CAFs in co-culture systems. |
| Pimonidazole HCl | Hypoxyprobe | Chemical probe for immunohistochemical detection of hypoxic regions in PDX tumors. |
| Anti-Human CD31 Antibody | Agilent Dako, Bio-Rad | Standard marker for identifying and quantifying human endothelial cells in PDX vasculature. |
| Luminex Multiplex Cytokine Panel | R&D Systems, Thermo Fisher | Profiles secretome from co-cultures to quantify stromal-tumor signaling (IL-6, VEGF, CCL2, etc.). |
| YAP/TAZ Antibody | Cell Signaling Tech, Santa Cruz | Detects nuclear translocation as a readout of mechanotransduction signaling. |
| Recombinant Hyaluronidase (PEGPH20-like) | Sigma-Aldrich, STEMCELL | Enzyme for experimental depletion of hyaluronan in ECM to study its role. |
Addressing Genetic Drift and Phenotypic Shift Across Passages
Patient-derived xenografts (PDXs) and organoids (PDOs) are indispensable tools in precision oncology, faithfully capturing the genomic and phenotypic heterogeneity of patient tumors. A core thesis underpinning their use is that these models provide a biologically relevant, patient-centric platform for investigating tumor biology, predicting therapeutic response, and advancing drug development. However, the fidelity of these models across extended in vitro and in vivo passages is not absolute. Genetic drift (accumulation of novel genomic alterations) and phenotypic shift (changes in morphology, marker expression, and drug response) threaten the long-term stability and clinical relevance of PDX and PDO lines. This guide details the mechanisms, monitoring strategies, and mitigation protocols essential for validating model fidelity within a rigorous research framework.
Mechanisms and Drivers of Instability
| Factor | Impact on PDX | Impact on PDO | Primary Consequence |
|---|---|---|---|
| Mouse Stromal Engraftment | High: Murine stroma replaces human tumor microenvironment (TME). | Not Applicable | Phenotypic shift; altered signaling from murine cytokines/growth factors. |
| Selection Pressure | High: Selection for clones adapted to murine TME or against human TME-dependent clones. | Moderate: Selection for clones adapted to specific culture Matrigel & media. | Genetic drift; loss of original tumor subpopulations. |
| Culture Conditions | Low (for tumor cells ex vivo). | Very High: Absence of physiological TME, non-physiological Matrigel, media composition. | Phenotypic shift; altered differentiation state & metabolism. |
| Passaging Method | Mechanical/ enzymatic disruption. | Enzymatic (e.g., TrypLE) vs. mechanical (e.g., chopping). | Bottlenecks that can drive genetic drift. |
| Cumulative Population Doublings | Moderate (slower than PDO). | Very High: Rapid expansion accelerates drift. | Accumulation of copy number variations (CNVs) and point mutations. |
Monitoring and Characterization Protocols
3.1. Genomic Fidelity Assessment
- Experiment: Short Tandem Repeat (STR) Profiling & SNP Genotyping.
- Protocol: Isolate DNA (DNeasy Blood & Tissue Kit) from early- and late-passage models and the original patient tumor (if available). Amplify a standardized panel of STR loci (e.g., 16 loci) and a genome-wide SNP array (e.g., Illumina Infinium). Compare profiles.
- Data Use: STR confirms human origin and identity, ruling out cross-contamination. SNP arrays quantify changes in allele frequency and detect large-scale CNVs indicative of clonal selection.
Experiment: Whole Exome Sequencing (WES) for Variant Allele Frequency (VAF) Tracking.
- Protocol: Perform WES (Illumina NovaSeq) on matched patient tumor, early-passage (P3-P5), and late-passage (P>10) samples. Align reads (BWA), call variants (GATK), and calculate VAF for driver mutations and a random subset of passenger mutations.
Quantitative Data:
Sample Median VAF of Key Driver Mutations Number of Private Novel SNVs (vs. Patient) Notes Patient Tumor (Primary) TP53: 45%, KRAS: 32% 0 (Baseline) Heterogeneous sample. PDX Passage 3 (F3) TP53: 78%, KRAS: 80% 12 Evidence of selection and mouse adaptation. PDX Passage 10 (F10) TP53: 95%, KRAS: 98% 45 Significant genetic drift; dominant clone emerged. PDO Passage 5 TP53: 70%, KRAS: 28% 25 Selection against KRAS-mutant clone in culture.
3.2. Phenotypic Fidelity Assessment
- Experiment: Immunohistochemistry (IHC) & Flow Cytometry for Marker Expression.
- Protocol: For PDXs: Fix, paraffin-embed, section. For PDOs: collect, make into a cytospin or micro-tissue array. Perform IHC for lineage markers (e.g., Cytokeratin), differentiation markers, and proliferation markers (Ki67). For flow cytometry, dissociate samples to single cells, stain with fluorescent-conjugated antibodies (e.g., CD44, CD133, EpCAM), and analyze.
- Data Use: Quantifies shifts in differentiation state and cancer stem cell populations.
Experiment: High-Throughput Drug Sensitivity & Resistance Testing (DSRT).
- Protocol: Plate dissociated PDOs or PDX-derived cells in 384-well plates. Treat with a library of 100+ oncology compounds across 5 concentrations (1 nM - 10 µM) for 72-96 hours. Assess viability via CellTiter-Glo. Calculate AUC (Area Under the curve) or IC50.
Quantitative Data:
Compound (Target) Patient-Derived Organoid IC50 (µM) Shift 5-Fluorouracil (Chemotherapy) Early Passage (P3): 1.2 ± 0.3 Baseline Late Passage (P15): 0.3 ± 0.1 Increased Sensitivity Olaparib (PARP) Early Passage (P3): 0.05 ± 0.01 Baseline Late Passage (P15): >10 Acquired Resistance
Mitigation Strategies and Best Practices
- Minimize Passages: Establish a central cryobank of low-passage stocks (PDX F3, PDO P3). Use for experiments directly upon revival; avoid continuous passaging.
- Orthogonal Authentication: Regularly authenticate models via STR, SNP fingerprinting, and Mycoplasma testing.
- Media Optimization: For PDOs, use defined media tailored to the tumor type, minimizing unknown selective factors. Conduct media component titration experiments.
- Cryopreservation of Intermediate Stocks: Freeze multiple vials at every other passage to create a "passage tree," allowing rollback to a more representative version.
- Co-culture Systems: For PDOs, incorporate relevant stromal cells (cancer-associated fibroblasts, immune cells) to preserve TME cues and reduce selective pressure.
- Early-Passage Use for Key Studies: Prioritize models below a validated passage threshold (e.g., PDX < F5, PDO < P10) for drug efficacy and biomarker studies.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Rationale |
|---|---|
| Matrigel, Growth Factor Reduced | Provides a 3D scaffold for PDO growth. The "Reduced" version minimizes unknown selective growth factor influences. |
| Defined Organoid Culture Media Kits | Tumor-type specific media (e.g., IntestiCult, Prostate Organoid Kit) provide consistent, defined conditions to limit drift. |
| Gentle Cell Dissociation Reagent | Enzymatic passaging reagents like TrypLE or dispase cause less damage than trypsin, reducing selection pressure. |
| PCR-Based Mycoplasma Detection Kit | Essential for routine screening; mycoplasma contamination drastically alters cell behavior and induces phenotypic shift. |
| Human/Mouse Species-Specific Antibodies | For flow cytometry/IHC to distinguish human tumor cells from murine stromal infiltration in PDXs (e.g., anti-HuNA, anti-MuNA). |
| CryoStor CS10 | Serum-free, defined cryopreservation medium superior to traditional DMSO/FBS mixes, improving post-thaw viability and recovery fidelity. |
Visualizations
Title: PDX/PDO Workflow with Fidelity Checkpoints
Title: Drivers of Model Drift and Shift
Best Practices for Biobanking, Annotation, and Data Management
Within the accelerating field of cancer research, patient-derived xenografts (PDXs) and organoids (PDOs) have emerged as transformative preclinical models that preserve the genetic, phenotypic, and functional heterogeneity of original tumors. The true translational power of these models is unlocked only through rigorous, standardized practices in biobanking, annotation, and data management. This whitepaper details the core technical best practices essential for generating high-fidelity, reproducible, and actionable research from PDX and PDO platforms.
Biobanking: From Patient to Preserved Sample
Effective biobanking is the cornerstone of reliable PDX/PDO research. The process must ensure virologic integrity and traceability.
Standard Operating Procedure for Tissue Acquisition and Processing
- Informed Consent & Ethical Compliance: All tissue collection must adhere to IRB-approved protocols with explicit consent for biobanking and downstream research.
- Ischemia Time Minimization: Target a cold ischemia time of <30 minutes. Document the exact time from devascularization to preservation.
- Macrodissection and Triaging: In a sterile, cold environment, dissect away necrotic and normal tissue. Split the sample for:
- Implantation (PDX: 3-5 mm³ fragments in cold Matrigel/PBS; PDO: mechanical/ enzymatic digestion).
- Cryopreservation (Snap-freezing in liquid N₂ for molecular analysis; viably freezing in cryoprotectant media).
- Histology (Fixation in 10% Neutral Buffered Formalin).
- Cryopreservation Protocol:
- Cell Suspension/Organoid Cryopreservation: Resuspend pellet in freezing medium (e.g., 90% FBS/10% DMSO or commercial organoid freezing medium). Cool at -1°C/min using a controlled-rate freezer or isopropanol chamber to -80°C before transfer to liquid nitrogen vapor phase.
Quantitative Data on Sample Quality
Table 1: Key Quality Metrics for Biobanked PDX/PDO Samples
| Metric | Target Specification | Measurement Method |
|---|---|---|
| Viability Post-Thaw | >70% for PDOs; Successful PDX engraftment from frozen fragment | Trypan blue exclusion; in vivo engraftment rate |
| RNA Integrity Number (RIN) | ≥7.0 for transcriptomic analysis | Bioanalyzer/TapeStation |
| Tumor Cellularity | ≥60% for genomic analysis | Pathologist review of H&E section |
| Mycoplasma Contamination | Negative | PCR-based or luminescence assay |
| Passage Number (PDX) | Documented; limit to <10 to minimize drift | Logbook and database tracking |
Diagram 1: Biobanking workflow for PDX and PDO models.
Comprehensive Annotation: The Metadata Backbone
Annotation transforms a biological sample into a research asset. FAIR (Findable, Accessible, Interoperable, Reusable) principles must guide data collection.
Minimum Information Framework
- Patient/Donor Level: Demographics, diagnosis (with histopathology report), treatment history, genomics (if available).
- Sample Level: Collection details (date, method, ischemia time), tumor cellularity, location.
- Model Generation Level: Engraftment site/method (PDX), culture matrix & media (PDO), passage number, splitting ratio.
- Characterization Level: Histology (IHC markers), genomic profile (WES/RNA-seq), STR fingerprinting for human cell ID, mycoplasma status.
Table 2: Essential Annotation Fields for PDX/PDO Models
| Tier | Field | Example Data Type | Critical for |
|---|---|---|---|
| Donor | Prior Therapies | Text List | Study resistance mechanisms |
| Sample | Tumor Cellularity (%) | Integer | Molecular analysis QC |
| Model | Engraftment Latency (PDX) | Days | Monitoring drift |
| Model | Culture Media Formula (PDO) | Text | Reproducibility |
| QC | STR Profile Match to Donor | Boolean (Y/N) | Identity confirmation |
| QC | Pathogen Screening Result | Text (Negative/Positive) | Contamination prevention |
Data Management and Integration
Robust data management systems are required to handle multi-omics, phenotypic, and experimental data.
Recommended System Architecture
- Laboratory Information Management System (LIMS): Tracks physical sample location, lineage (passaging), and aliquots.
- Electronic Lab Notebook (ELN): Documents experimental protocols and results.
- Metadata Repository: A centralized, structured database (e.g., based on MIABIS or BRISQ standards) stores all sample annotation.
- Analysis Data Warehouse: Securely houses raw and processed genomic, imaging, and drug response data.
Experimental Protocol: High-Throughput Drug Screening in PDOs
- Organoid Harvest & Dissociation: Mechanically and enzymatically dissociate a validated PDO culture into single cells/small clusters.
- Cell Counting & Viability Assessment: Use automated cell counter with trypan blue. Adjust concentration to target seeding density.
- Seeding: Plate 5-10,000 cells/well in 5μL BME/Matrigel droplets in 384-well plates. Centrifuge, polymerize (37°C, 30 min).
- Drug Addition: Using a D300e digital dispenser, add compounds in 7-point, 1:3 serial dilution in 50μL complete media. Include DMSO controls.
- Incubation: Culture for 5-7 days, with media refresh at day 3-4.
- Viability Readout: Add CellTiter-Glo 3D reagent, shake, incubate, and measure luminescence.
- Data Analysis: Normalize to controls, fit dose-response curves (e.g., using R
drcpackage), calculate IC₅₀/ AUC values.
Diagram 2: PDO high-throughput drug screening workflow.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for PDX/PDO Biobanking and Screening
| Item | Function/Application | Key Consideration |
|---|---|---|
| Basement Membrane Extract (BME) | 3D extracellular matrix for PDO embedding & growth. | Lot-to-lot variability; keep on ice to prevent polymerization. |
| Cryoprotectant Medium | Viable freezing of cell/organoid suspensions. | Use DMSO-containing medium or specialized serum-free, animal-free commercial options. |
| Digestion Enzymes | Tissue dissociation for PDO generation and passaging. | Combine collagenase, dispase, or gentle, tumor-specific cocktails. |
| Stem Cell Factor-enriched Media | Supports growth of tumor stem/progenitor cells in PDOs. | Often requires WNT/R-spondin, Noggin, EGF; formulations are cancer-type specific. |
| CellTiter-Glo 3D | Luminescent ATP assay for 3D culture viability quantification. | Requires extended shaking/incubation for reagent penetration into organoids. |
| STR Profiling Kit | Human cell line authentication and confirmation of donor match. | Mandatory QC step to exclude cross-contamination or misidentification. |
| Mycoplasma Detection Kit | Routine screening for bacterial contamination. | Use highly sensitive PCR-based methods monthly. |
| Digital Dispenser (e.g., D300e) | Non-contact, precise delivery of drug compounds for screening. | Enables high-density plate formats and complex dose-response designs. |
The integration of meticulous biobanking, exhaustive annotation, and rigorous data management creates a robust foundation for PDX and PDO research. By adhering to these technical best practices, research consortia and individual labs can ensure the quality, reproducibility, and interoperability of their models, thereby accelerating the translation of discoveries from these high-fidelity systems into meaningful clinical advances in oncology.
PDX vs. PDO: A Head-to-Head Comparison of Strengths, Limitations, and Ideal Use Cases
The integration of patient-derived xenografts (PDXs) and patient-derived organoids (PDOs) has revolutionized preclinical cancer research by providing more clinically relevant models. This guide provides a direct, technical comparison of these two platforms across critical operational parameters—throughput, cost, timeline, and scalability—to inform experimental design and resource allocation in research and drug development.
Quantitative Comparison of Core Parameters
Table 1: Direct Comparison of PDX vs. PDO Platforms
| Parameter | Patient-Derived Xenograft (PDX) | Patient-Derived Organoid (PDO) |
|---|---|---|
| Throughput (Experimental) | Low to Moderate. Limited by mouse housing, engraftment time, and serial passaging. Typical cohort size: 5-10 mice per group. | High. Enabled by in vitro culture in multi-well plates. Dozens to hundreds of replicates/conditions per experiment. |
| Establishment Success Rate | ~30-70%, highly variable by cancer type. | ~50-90%, generally higher for epithelial cancers. |
| Time to Initial Experiment (Latency) | Long (3-8 months). Includes engraftment (2-6 months) and expansion in mice (P1). | Short (2-8 weeks). Includes initial organoid formation (1-4 weeks) and first expansion passage. |
| Cost per Model/Line (Initial) | High ($3,000 - $10,000+). Includes host mice, surgery/imaging, long-term husbandry, and biosafety costs. | Moderate ($500 - $2,500). Includes specialized matrices, growth factors, and culture reagents. |
| Cost per Drug Screening Assay | Very High ($5,000 - $15,000+). Driven by mouse numbers, compound requirements, and lengthy in vivo monitoring. | Low ($100 - $1,000). Scalable in plates with minimal compound volumes. |
| Scalability (Parallel Testing) | Poor. Difficult to scale due to animal logistics, cost, and ethical constraints. | Excellent. Amenable to high-throughput automation, liquid handlers, and high-content imaging. |
| Genetic & Histopathological Stability | High over early passages (<5), but can drift due to murine stromal selection. | High, but culture conditions may select for specific clones; requires monitoring. |
| Tumor Microenvironment (TME) | Complex, human-murine hybrid. Retains human cancer cells with recruited murine stroma, vasculature, and immune system (in humanized models). | Simplified, mostly epithelial. Primarily human cancer cells with potential for co-culture (CAFs, immune cells) in engineered systems. |
Detailed Experimental Protocols
Protocol 1: PDX Establishment & Drug Efficacy Study
Aim: To establish a PDX model from a human tumor sample and evaluate in vivo drug response.
Methodology:
- Tumor Implantation: Fresh tumor fragments (1-3 mm³) or dissociated cells (Matrigel suspended) are surgically implanted into immunocompromised host mice (e.g., NSG) subcutaneously, orthotopically, or under the renal capsule.
- Engraftment Monitoring: Tumor growth is monitored weekly via caliper measurements. Engraftment is considered successful upon reaching a volume of ~500 mm³ (F0 generation).
- Passage & Expansion: The primary (F0) tumor is harvested, dissociated, and re-implanted into new mice to generate P1 (F1) stocks. A cryobank is established.
- Drug Efficacy Study:
- Randomization: When P2 tumors reach ~150-200 mm³, mice are randomized into treatment and control groups (n=5-10).
- Dosing: Test compound or vehicle is administered via the intended route (oral, IP, IV) at a defined schedule.
- Endpoint Analysis: Tumor volume and mouse weight are tracked 2-3 times weekly. At study end, tumors are harvested for downstream analysis (IHC, RNA-seq, pharmacodynamics).
Key Reagents:
- Basement Membrane Matrix (e.g., Matrigel): Provides structural support for cell implantation.
- Immunodeficient Mice (NSG): Hosts that allow engraftment of human tissue.
- Analgesics & Anesthetics (Buprenorphine, Isoflurane): For ethical post-operative care.
Protocol 2: PDO Generation & High-Throughput Drug Screening
Aim: To establish a PDO biobank from patient tissue and perform a dose-response drug screen.
Methodology:
- Tissue Processing: Fresh tumor tissue is minced and enzymatically digested (Collagenase/Dispase) to a single-cell/small-cluster suspension.
- Embedding & Culture: Cells are mixed with a reduced-growth factor Basement Membrane Extract (BME) and plated as droplets in a pre-warmed culture plate. After BME polymerization, a defined organoid growth medium (containing Wnt3A, R-spondin, Noggin, EGF, etc.) is overlaid.
- Expansion & Passaging: Organoids are grown for 7-14 days, then mechanically/ enzymatically dissociated and re-embedded in fresh BME for passaging.
- High-Throughput Screening:
- Seeding: Dissociated organoids are seeded into 384-well assay plates in BME.
- Drug Treatment: After 3-5 days, compounds are added via acoustic liquid dispensing in a dose-response matrix (e.g., 8-point, 1:3 serial dilution).
- Viability Readout: After 5-7 days of treatment, cell viability is assessed via ATP-based luminescence (CellTiter-Glo 3D).
- Data Analysis: Dose-response curves are fitted, and IC50/Area-Under-Curve values are calculated for each compound-line pair.
Key Reagents:
- BME, Type 2 / Cultrex: Provides the essential 3D extracellular matrix scaffold.
- Advanced DMEM/F12: Base medium for organoid culture.
- Recombinant Growth Factors (Wnt3A, R-spondin-1, Noggin): Essential for stem cell maintenance and proliferation in epithelial organoids.
- Rho-Kinase Inhibitor (Y-27632): Prevents anoikis during passaging.
- CellTiter-Glo 3D: Luminescent assay optimized for 3D culture viability measurement.
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for PDX and PDO Research
| Reagent Category | Specific Example(s) | Function in Context |
|---|---|---|
| Extracellular Matrix | Matrigel (Corning), Cultrex BME (R&D Systems) | Provides a 3D scaffold mimicking the basement membrane; essential for engraftment (PDX) and organoid growth (PDO). |
| Specialized Media | PDO: IntestiCult (StemCell Tech), PDX: DMEM/F12 with serum. | Formulated with essential nutrients and, for PDOs, specific growth factors to support the stem cell niche. |
| Growth Factors | Recombinant Human EGF, Wnt3A, R-spondin-1, Noggin (PeproTech, R&D) | Critical signaling molecules for maintaining stemness and driving proliferation in PDO cultures. |
| Dissociation Enzymes | Collagenase IV, Dispase, TrypLE Express (Thermo Fisher) | Gently dissociate tissue into single cells/small clusters for PDO generation or PDX passaging. |
| Apoptosis Inhibitor | Y-27632 (ROCK inhibitor) | Specifically used in PDO culture to inhibit dissociation-induced cell death during passaging. |
| Viability Assay | CellTiter-Glo 3D (Promega) | Optimized luminescent assay for measuring ATP in 3D structures like organoids. |
| Host Strain | NOD-scid IL2Rγ[null] (NSG) mice (The Jackson Lab) | Gold-standard immunodeficient host for PDX engraftment due to lack of adaptive immunity. |
Visualization of Workflows and Signaling
PDX Establishment & In Vivo Study Workflow
PDO Biobanking & High-Throughput Screening Workflow
Core Signaling Pathways in Epithelial PDO Culture
Within the broader thesis on Introduction to Patient-Derived Xenografts (PDXs) and Organoids in Cancer Research, a critical pillar is the rigorous validation of these advanced preclinical models. Their utility in translational research, drug discovery, and personalized medicine hinges on their fidelity to the original patient tumor. This guide provides an in-depth technical analysis of the core concept of model fidelity, defined by the concordance between a model (PDX or organoid) and its tumor of origin across three fundamental molecular and phenotypic layers: genomic, transcriptomic, and histopathological.
Defining Fidelity and Concordance
Fidelity refers to the degree to which a model retains the defining characteristics of the source tumor after establishment and propagation. Concordance is the measurable similarity between the model and the tumor across specific data types. High multi-modal concordance increases confidence that therapeutic responses observed in the model will predict clinical outcomes.
The following tables summarize typical concordance metrics reported in recent literature for early-passage PDXs and matched organoids.
Table 1: Genomic Concordance Metrics (SNVs/Indels/Copy Number Variations)
| Model Type | Passage | Median SNV Concordance | Key CNV Retention | Notable Drift |
|---|---|---|---|---|
| PDX (P0-P3) | Early | 85-95% | Major driver amplifications/deletions | Murine stromal overgrowth in later passages |
| Organoid (P5-P15) | Early-Mid | >90% | Tumor-specific copy number profile | Culture adaptation (e.g., APC truncations in CRC) |
| Analysis Method | Platform | Typical Coverage | Key Metric | Threshold |
| Whole Exome Sequencing | Illumina NovaSeq | >100x | Variant Allele Frequency Correlation | R² > 0.85 |
| SNP Array/CNV Kit | Affymetrix/OncoScan | N/A | Copy Number Profile Pearson Correlation | r > 0.8 |
Table 2: Transcriptomic and Histopathological Concordance
| Analysis Layer | Measurement Tool | Typical Concordance Range (High-Fidelity Models) | Primary Cause of Discordance |
|---|---|---|---|
| Transcriptomic | RNA-Seq, Gene Expression Microarray | 70-90% (Spearman correlation) | Stromal contamination (PDX), culture media selection (organoids) |
| Pathway Activity | GSEA, Single-Sample Signatures | Significant overlap (FDR < 0.25) | Microenvironmental differences |
| Histopathological | H&E Staining, IHC | Morphology: High; IHC marker score: 80-95% | Loss of tumor-infiltrating lymphocytes, architectural simplification |
Detailed Experimental Protocols for Assessing Fidelity
Protocol 1: Genomic Concordance Analysis via Whole Exome Sequencing (WES)
Objective: To identify and compare somatic variants and copy number alterations between patient tumor (PT), matched PDX, and/or organoid models.
Materials:
- DNA from PT (FFPE or frozen), PDX (snap-frozen, murine stroma depleted), and organoids.
- Qubit dsDNA HS Assay Kit, Agilent SureSelect XT Human All Exon v8.
- Illumina sequencing platform (e.g., NovaSeq 6000).
Procedure:
- DNA Extraction & QC: Extract high-quality DNA (≥100ng, integrity number ≥7). Use magnetic bead-based murine DNA depletion kits for PDX samples (e.g., NuGEN NuProbe Mouse Depletion).
- Library Preparation & Enrichment: Perform shearing, end-repair, adapter ligation, and PCR amplification per kit instructions. Hybridize with biotinylated baits for human exome capture.
- Sequencing: Sequence to a minimum mean coverage of 100x for human-derived reads.
- Bioinformatic Analysis:
- Alignment: Map reads to the human reference genome (hg38) using BWA-MEM.
- Variant Calling: Call somatic SNVs/Indels with MuTect2 (GATK) using the PT as "tumor" and a matched normal (patient blood) as "reference." Call variants in models against the same normal.
- Concordance Calculation: Calculate the overlap of somatic variants (VAF > 0.1) between PT and model. Generate a scatter plot of Variant Allele Frequencies for shared variants.
- CNV Analysis: Use control-FREEC or GATK gCNV to generate copy number profiles. Calculate Pearson correlation between log2 ratio profiles of PT and model.
Protocol 2: Transcriptomic Profiling and Concordance Assessment
Objective: To compare global gene expression patterns and pathway activities.
Materials: TRIzol or RNEasy Kit, Poly-A selection beads, Illumina Stranded mRNA Prep.
Procedure:
- RNA Extraction & QC: Extract total RNA. Ensure RIN ≥ 8.5.
- Library Prep & Sequencing: Perform poly-A selection, cDNA synthesis, adapter ligation, and sequencing (≥30M paired-end reads).
- Bioinformatic Analysis:
- Quantification: Align reads with STAR and quantify gene counts using featureCounts.
- Deconvolution (for PDX): Use tools like
xenograftorDeMixTto estimate human vs. mouse RNA contribution and infer pure human tumor expression. - Concordance Metrics: Perform Spearman correlation on normalized (TPM, voom) expression of the top 5000 most variable genes. Conduct Gene Set Enrichment Analysis (GSEA) to compare enrichment scores of hallmark pathways.
Protocol 3: Histopathological and Immunophenotypic Analysis
Objective: To assess preservation of tumor morphology and protein marker expression.
Materials: Formalin-fixed, paraffin-embedded (FFPE) blocks, H&E stain, automated IHC stainer, validated primary antibodies.
Procedure:
- Sectioning & Staining: Cut 4-5μm sections. Perform standard H&E staining.
- Immunohistochemistry (IHC): Perform antigen retrieval, block, incubate with primary antibodies (e.g., Cytokeratin, ER, HER2, Ki-67), and develop with DAB.
- Scoring & Analysis:
- Morphology: A blinded pathologist reviews H&E slides for architectural patterns, cell morphology, and stromal components.
- IHC Quantification: Use semi-quantitative scoring (e.g., H-score for ER/PR: intensity 0-3 x % positive cells) or digital image analysis (e.g., QuPath) for objective quantification.
- Concordance: Report percentage agreement for categorical markers (positive/negative) and correlation coefficients for continuous scores.
Visualizing Workflows and Biological Relationships
Title: Multi-Omic Fidelity Assessment Workflow
Title: Retention of Key Genomic Drivers and Pathway Activity
The Scientist's Toolkit: Key Research Reagent Solutions
Table 3: Essential Reagents for Fidelity Analysis Experiments
| Reagent/Material | Supplier Examples | Primary Function in Fidelity Analysis |
|---|---|---|
| Murine DNA Depletion Kit | NuGEN (NuProbe), IDT (xGen) | Selectively depletes mouse-derived genomic DNA from PDX samples, enriching human tumor DNA for sequencing. |
| Human/Mouse Species-Specific Panels | NanoString (PanCancer IO 360 Mouse), ArcherDx (Immunoverse) | Enable quantitation of species-specific RNA transcripts in PDX for deconvolution and accurate human transcriptome analysis. |
| Matrigel, Cultrex BME | Corning, Bio-Techne | Basement membrane extract for 3D organoid culture, supporting architecture resembling the original tumor. |
| Tumor Dissociation Kits | Miltenyi Biotec (Tumor Dissociation Kit), STEMCELL Tech. | Gentle enzymatic dissociation of tumor tissue into single cells for initiating PDX/organoid models. |
| FFPE RNA/DNA Extraction Kits | Qiagen (AllPrep), Thermo Fisher (RecoverAll) | Isolate nucleic acids from archived patient FFPE samples for direct comparison with model material. |
| Multiplex IHC/IF Kits | Akoya (PhenoCycler), Standard Biotools (Codex) | Simultaneously profile 30+ protein markers on a single tissue section, enabling deep spatial phenotyping concordance. |
| Targeted Sequencing Panels | Illumina (TruSight Oncology 500), Tempus (xT) | Profile hundreds of cancer genes for SNVs, indels, CNVs, and fusions in PT and models for cost-effective concordance checks. |
Achieving and validating high genomic, transcriptomic, and histopathological concordance is non-negotiable for leveraging PDX and organoid models in rigorous translational research. The protocols and metrics outlined here provide a framework for systematic fidelity analysis. As these models increasingly guide preclinical drug development and personalized treatment strategies, continuous multi-modal benchmarking against the patient tumor remains the cornerstone of their scientific and predictive value.
Patient-derived xenografts (PDXs) have emerged as a cornerstone of translational cancer research, offering a model system that bridges the gap between in vitro studies and human clinical trials. Within the broader thesis of integrating PDXs and organoids into cancer research, PDXs provide unique and critical advantages in three key areas: recapitulating in vivo physiology, maintaining an intact tumor microenvironment (TME), and enabling the study of metastasis. This guide details the technical foundations of these strengths.
In Vivo Physiology
PDX models are established by implanting fresh patient tumor tissue, either subcutaneously or orthotopically, into immunocompromised mice (e.g., NSG, NOG). This process preserves the original tumor's histopathological architecture, genetic heterogeneity, and stromal components through early passages. The in vivo setting allows for the study of tumor growth kinetics, pharmacodynamic responses, and host-organ interactions that are impossible to replicate in a dish.
Key Quantitative Data on PDX Physiology Fidelity
| Physiological Parameter | PDX Model Retention Rate (%) | Notes / Key Study Findings |
|---|---|---|
| Genetic Heterogeneity | 85-95% | Whole-exome sequencing shows high concordance of somatic mutations between patient tumor and early-passage PDX (passage 3-5). |
| Transcriptomic Profile | 80-90% | RNA-seq analyses reveal conserved gene expression signatures, though murine stromal infiltration increases with passaging. |
| Histopathological Architecture | >90% | H&E staining demonstrates preservation of original tumor morphology and grading in early passages. |
| Drug Response Correlation | 70-85% | Retrospective studies show PDX response to chemotherapies often correlates with the patient's clinical outcome. |
Experimental Protocol: Establishing a Subcutaneous PDX Model
- Tumor Acquisition: Obtain fresh tumor tissue from surgical resection or biopsy in sterile, cold transport media.
- Processing: Mince tissue into ~1-3 mm³ fragments in a biosafety cabinet. Optional enzymatic digestion (Collagenase/Hyaluronidase) can be performed for cell suspension implantation.
- Mouse Preparation: Anesthetize an immunodeficient mouse (e.g., 8-10 week old NSG).
- Implantation: Using a trocar, implant one fragment subcutaneously into the flank. For cell suspensions, mix with Matrigel and inject subcutaneously.
- Monitoring: Monitor tumor growth via caliper measurements. Upon reaching ~1000-1500 mm³, harvest for serial passaging or analysis.
Intact Microenvironment
While the human immune system is largely absent in standard PDX models, the implanted tumor brings its native human cancer-associated fibroblasts (CAFs), endothelial cells, and extracellular matrix (ECM). The mouse host then contributes a living, responsive murine stroma, including blood vessels that undergo angiogenesis to support the tumor. This intact, albeit chimeric, microenvironment is crucial for studying stroma-mediated drug resistance and tumor biology.
Research Reagent Solutions for TME Analysis
| Reagent / Material | Function / Application |
|---|---|
| Anti-Human HLA-A,B,C Antibody | IHC staining to distinguish human tumor cells from murine stroma. |
| Anti-Mouse CD31 Antibody | IHC staining to visualize mouse-derived endothelial cells and vasculature. |
| Anti-αSMA Antibody | Labels cancer-associated fibroblasts (CAFs) in the stroma. |
| Masson's Trichrome Stain | Histological stain to visualize collagen and ECM composition. |
| Species-Specific FISH Probes | Fluorescent in situ hybridization probes to identify human vs. mouse genomic DNA in sections. |
| Luminex Cytokine Assay (Mouse) | Quantifies mouse-derived cytokines and growth factors in the TME from tumor homogenates. |
Metastasis Studies
Orthotopic implantation of PDXs—placing breast cancer tissue into the mouse mammary fat pad or colorectal cancer cells into the cecum—provides the correct anatomical and hormonal context for studying metastatic spread. This enables the investigation of the entire metastatic cascade: local invasion, intravasation, circulation, extravasation, and colonization of distant organs (e.g., lung, liver, bone).
Experimental Protocol: Orthotopic Implantation for Metastasis Tracking
- Cell Preparation: Generate a luciferase-tagged single-cell suspension from a donor PDX tumor.
- Mouse Preparation: Anesthetize and sterilize the surgical site on an NSG mouse.
- Orthotopic Injection: For mammary fat pad: make a small incision, expose the fat pad, and inject cells. For intrasplenic or intrahepatic injections: perform a laparotomy.
- In Vivo Imaging: Weekly, inject mouse with D-luciferin substrate and image using an IVIS Spectrum in vivo imaging system to quantify primary tumor growth and metastatic spread via bioluminescence.
- Endpoint Analysis: Harvest primary tumor and suspected metastatic organs for ex vivo bioluminescence imaging and histopathological confirmation (H&E staining).
Key Quantitative Metastasis Data from PDX Studies
| Metastatic Parameter | Typical Measurement in PDX Models | Common Assay / Technology |
|---|---|---|
| Metastatic Incidence | 30-70% (varies by tumor type) | Histological scoring of organs from orthotopic models. |
| Metastatic Burden | Bioluminescence Radiance (p/s/cm²/sr) | In vivo imaging system (IVIS) quantification. |
| Circulating Tumor Cells (CTCs) | 10-1000 cells/mL blood | Flow cytometry (human CD298+ mouse CD45-) or microfluidics. |
| Time to Metastasis | 4-16 weeks post-implantation | Weekly monitoring via imaging. |
| Organotropism Pattern | Reflects patient disease (e.g., lung, liver) | Ex vivo organ imaging and histology. |
Within the integrated paradigm of PDXs and organoids, PDXs provide the indispensable in vivo platform that captures physiological complexity. Their strengths in modeling intact, dynamic microenvironments and the multi-step process of metastasis make them unparalleled for preclinical drug efficacy testing, biomarker discovery, and investigating fundamental mechanisms of tumor progression. These models are most powerful when used in concert with in vitro organoid systems, which offer complementary strengths in high-throughput screening and genetic manipulation.
The evolution of preclinical cancer models from cell lines to patient-derived xenografts (PDXs) and now to patient-derived organoids (PDOs) represents a paradigm shift towards greater clinical fidelity. While PDXs maintain a stromal microenvironment and are invaluable for in vivo studies, they are costly, slow to establish, and less amenable to scalable experimentation. PDOs, three-dimensional ex vivo cultures derived directly from patient tissue, bridge this gap. They retain key genetic and phenotypic characteristics of the original tumor while offering the manipulability and scalability of traditional cell culture. This whitepaper details the core strengths of PDOs—high-throughput screening potential, precise genetic manipulation, and application in personalized therapeutic screens—positioning them as a complementary and indispensable tool alongside PDXs in modern translational oncology research.
High-Throughput Screening (HTS) Potential
PDOs can be established, expanded, and banked, enabling their use in large-scale drug discovery campaigns. Their growth in multi-well plates (96-, 384-, 1536-well formats) allows for the simultaneous testing of hundreds to thousands of compounds.
Key Quantitative Data on PDO HTS:
| Metric | Typical Range/Value | Comparison to 2D Lines | Notes |
|---|---|---|---|
| Establishment Success Rate | 50-90% (varies by tumor type) | Higher than PDX (30-70%) | Gastrointestinal cancers show highest rates. |
| Expansion Potential | 10^4 - 10^6 cells per well in 2-4 weeks | Slower than 2D, but sufficient for HTS | Requires optimized, often expensive, niche-specific media. |
| Assay Miniaturization | 10-20 µL final volume in 1536-well plates | Comparable | Viability readouts (CellTiter-Glo 3D) are standard. |
| Screening Throughput | 10,000+ compounds per week per system | Similar throughput, but with superior biological relevance. | Z'-factors often >0.5, indicating robust assay quality. |
| Drug Response Data Correlation with Clinical Outcome | 80-95% (Predictive Positive Value) | Significantly higher than 2D lines (<50%) | Based on retrospective studies in colorectal, pancreatic cancers. |
Experimental Protocol: High-Throughput Drug Screening on PDOs
- PDO Preparation: Mechanically or enzymatically dissociate mature PDOs into single cells or small clusters (<50 cells).
- Seeding: Using an automated liquid handler, plate cell suspension into ultra-low attachment spheroid microplates (e.g., Corning 384-well). Seed at an optimized density (e.g., 500-2000 cells/well).
- Recovery: Centrifuge plates briefly (100-300 x g, 1 min) to aggregate cells at the well bottom. Incubate for 3-5 days to allow re-formation of micro-organoids.
- Compound Addition: Pin-transfer or acoustically dispense compound libraries (nL volumes) from DMSO stocks. Include positive (e.g., 10µM Staurosporine) and negative (DMSO only) controls.
- Incubation: Incubate plates for 5-7 days, with media change at day 3-4 if necessary.
- Viability Assay: Add an equal volume of CellTiter-Glo 3D Reagent, shake orbially for 5 min, incubate for 25 min in the dark, and measure luminescence.
- Data Analysis: Normalize luminescence to controls. Calculate % viability and IC50/GR50 values using curve-fitting software (e.g., GraphPad Prism, Dotmatics).
High-Throughput Drug Screening with PDOs
Genetic Manipulation in PDOs
The capacity for precise genetic engineering allows functional genomics and isogenic modeling within a clinically relevant system. CRISPR-Cas9 is the primary tool for knockout, knock-in, and gene editing in PDOs.
Experimental Protocol: CRISPR-Cas9 Knockout in PDOs
- Guide RNA (gRNA) Design: Design 2-3 gRNAs targeting the gene of interest using online tools (e.g., Broad Institute's GPP Portal). Clone into a lentiviral expression vector (e.g., lentiCRISPRv2).
- Lentivirus Production: Co-transfect HEK293T cells with the gRNA vector, psPAX2 (packaging), and pMD2.G (envelope) plasmids using PEI transfection reagent. Harvest virus-containing supernatant at 48 and 72 hours.
- PDO Transduction: Dissociate PDOs to single cells. Spinoculate cells with lentiviral supernatant and polybrene (8µg/mL) at 1000 x g for 1 hour at 32°C. Plate cells in Matrigel.
- Selection & Expansion: After 48 hours, add appropriate antibiotic (e.g., Puromycin, Blasticidin) for 5-7 days to select transduced cells. Expand surviving organoids.
- Validation: Perform genomic DNA extraction, PCR amplify the target region, and sequence via TIDE analysis or next-generation sequencing to quantify editing efficiency. Validate protein loss via western blot.
The Scientist's Toolkit: Key Reagents for PDO Genetic Manipulation
| Reagent/Material | Function | Example Product |
|---|---|---|
| Lentiviral CRISPR Vector | Delivers Cas9 and gRNA for stable genomic integration. | lentiCRISPRv2 (Addgene #52961) |
| Packaging Plasmids | Required for production of replication-incompetent lentivirus. | psPAX2 (Addgene #12260), pMD2.G (Addgene #12259) |
| Transfection Reagent | Facilitates plasmid DNA delivery into producer cells. | Polyethylenimine (PEI Max) |
| Polybrene | A cationic polymer that enhances viral transduction efficiency. | Hexadimethrine bromide |
| Basement Membrane Matrix | Provides 3D scaffold for PDO growth post-transduction. | Corning Matrigel, GFR |
| Selection Antibiotic | Selects for cells successfully transduced with the resistance gene-bearing vector. | Puromycin, Blasticidin S |
CRISPR-Cas9 Gene Editing in PDOs
Personalized Therapeutic Screens
PDOs serve as a patient avatar for functional precision medicine. Co-clinical trials, where PDOs are derived and tested in parallel with a patient's treatment, can predict therapeutic response.
Key Quantitative Data on Personalized PDO Screens:
| Study Parameter | Reported Outcome | Clinical Implication |
|---|---|---|
| Turnaround Time (Biopsy to Data) | 3-6 weeks | Feasible for informing later lines of therapy. |
| Positive Predictive Value (PPV) | 85-100% | High confidence that a drug effective in PDO will work in the patient. |
| Negative Predictive Value (NPV) | 80-95% | High confidence that a drug ineffective in PDO will not work in the patient. |
| Success Rate in Guiding Therapy (Observational Studies) | ~70% | Demonstrates potential clinical utility. |
| Cost per PDO Screen (Compound Library) | $3,000 - $10,000 | Comparable to genomic sequencing; cost-effective if prevents ineffective therapy. |
Experimental Protocol: Co-Clinical Trial with PDOs
- Biopsy & PDO Generation: Obtain tumor tissue (surgical resection or biopsy) under IRB approval. Process immediately to establish PDO cultures in tumor-type specific media.
- Ex Vivo Drug Testing: Once sufficient biomass is expanded (P2-P3), screen against a panel of clinically relevant agents (standard-of-care, targeted therapies, investigational drugs). Use the HTS protocol above in a 96- or 384-well format.
- Data Integration & Analysis: Generate a drug response profile (e.g., waterfall plot of % viability). Integrate with the patient's genomic and transcriptomic data to identify mechanistic correlates of response/resistance.
- Clinical Reporting: A multidisciplinary tumor board reviews PDO data alongside clinical and molecular data. A report is generated for the treating oncologist, highlighting ex vivo sensitive and resistant agents.
- Clinical Correlation: The patient's subsequent treatment response is monitored and formally compared to the PDO prediction.
Co-Clinical Trial Workflow Using PDOs
Patient-derived organoids embody a transformative technology that addresses critical limitations of both traditional cell lines and PDXs. Their strengths—high-throughput potential for scalable drug discovery, precise genetic manipulation for mechanistic studies, and direct application in personalized therapeutic screens for functional precision medicine—establish them as a cornerstone of modern cancer research. When integrated into a holistic preclinical strategy alongside in vivo PDX models, PDOs significantly enhance the predictive power, efficiency, and clinical relevance of translational oncology, accelerating the path from bench to bedside.
Within the thesis on "Introduction to patient-derived xenografts and organoids in cancer research," selecting the appropriate preclinical model is not a mere preliminary step but a critical determinant of experimental validity and translational relevance. Patient-derived xenografts (PDXs) and patient-derived organoids (PDOs) have revolutionized oncology research by preserving the genetic and phenotypic heterogeneity of human tumors. However, their distinct biological and technical characteristics necessitate a deliberate alignment of model capabilities with specific research questions. This guide provides a technical framework for this alignment, underpinned by current data and protocols.
Comparative Analysis of Core Model Systems
The following table synthesizes quantitative and qualitative data from recent literature (searched 2023-2024) comparing PDX and PDO models across key parameters.
Table 1: Comparative Capabilities of PDX vs. PDO Models
| Parameter | Patient-Derived Xenografts (PDX) | Patient-Derived Organoids (PDOs) |
|---|---|---|
| Establishment Success Rate | 20-40% (varies by tumor type) | 50-80% (higher for epithelial cancers) |
| Time to Experimental Readiness | 4-12 months (incl. engraftment & expansion) | 2-8 weeks |
| Stromal/Immune Microenvironment | Murine stroma; lacks human immune system (unless humanized) | Primarily epithelial; can be co-cultured with fibroblasts/immune cells |
| Genetic Stability | High over early passages (<5); drift possible long-term | Generally high in defined culture conditions |
| Throughput Potential | Low to medium (in vivo constraints) | High (in vitro, 96/384-well compatible) |
| Cost per Model/Line | High (~$5k - $15k) | Moderate (~$1k - $5k) |
| Ideal Application | In vivo therapy response, metastasis, pharmacokinetics/pharmacodynamics (PK/PD) | High-throughput drug screening, mechanistic studies, personalized therapy guides |
Aligning Research Questions with Model Capabilities
Research Question 1: "What is the in vivo efficacy and optimal dosing schedule of a novel compound?"
- Recommended Model: PDX. This question requires an intact physiology for drug absorption, distribution, metabolism, and excretion (ADME).
- Key Experimental Protocol (PDX Treatment Study):
- PDX Expansion: Expand tumor fragment in NOD/SCID or NSG mice until ~200-400 mm³.
- Randomization & Cohorting: Randomize tumor-bearing mice into control and treatment groups (n≥5/group). Match starting tumor volumes across groups.
- Dosing: Administer compound via planned clinical route (e.g., oral gavage, IP, IV). Control groups receive vehicle.
- Monitoring: Measure tumor volumes (via caliper) and mouse weights 2-3 times weekly. Calculate tumor volume: (Length x Width²)/2.
- Endpoint Analysis: At study endpoint, harvest tumors. Perform weighing, formalin-fixation/paraffin-embedding (FFPE) for histology (H&E, IHC), and snap-freezing for molecular analysis (RNA/DNA extraction).
- Statistical Analysis: Compare tumor growth curves (e.g., repeated measures ANOVA) and final tumor weights/volumes.
Research Question 2: "Which drug, from a library of 100 candidates, shows the highest potency against a specific cancer subtype?"
- Recommended Model: PDO. This question demands high-throughput scalability and quantitative readouts.
- Key Experimental Protocol (PDO High-Throughput Drug Screen):
- PDO Preparation: Culture PDOs in basement membrane extract (e.g., Matrigel). Dissociate into single cells or small clusters using enzymatic digestion (e.g., TrypLE).
- Seeding: Seed cells into 384-well plates pre-coated with a thin layer of Matrigel in defined growth medium.
- Drug Treatment: After 24-48h, add compound library via automated liquid handler. Include DMSO vehicle and reference control (e.g., cisplatin) on each plate.
- Viability Assay: Incubate for 5-7 days. Add cell viability reagent (e.g., CellTiter-Glo 3D). Measure luminescence on a plate reader.
- Data Analysis: Normalize luminescence to vehicle control (100% viability) and reference control (0% viability). Calculate dose-response curves and IC₅₀ values for each compound.
Signaling Pathway & Workflow Visualizations
Diagram 1: PDX Establishment and Study Workflow
Diagram 2: PDO Culture and Expansion Workflow
Diagram 3: Core PI3K/AKT/mTOR Signaling Pathway
The Scientist's Toolkit: Essential Research Reagent Solutions
Table 2: Key Reagents for PDX and PDO Research
| Reagent/Material | Primary Function | Application Note |
|---|---|---|
| Basement Membrane Extract (Matrigel) | Provides 3D extracellular matrix scaffold for cell growth and polarization. | Essential for PDO culture. Use growth factor-reduced for defined conditions. Keep on ice. |
| Advanced DMEM/F-12 Medium | Base nutrient medium for organoid cultures. | Formulated for low serum conditions; basis for specialized supplements. |
| NICHE Factors (Wnt3a, R-spondin, Noggin) | Maintain stemness and support growth of epithelial stem cells. | Critical for gastrointestinal, hepatic, and other PDOs. Used as conditioned media or recombinant proteins. |
| Y-27632 (ROCK inhibitor) | Inhibits anoikis (cell death after detachment). | Significantly improves viability during PDO passaging and thawing. |
| Collagenase/Hyaluronidase | Enzymatic digestion of tumor tissue to release cell clusters. | Used in initial processing of patient samples for both PDX and PDO. |
| NOD/SCID/NSG Mice | Immunodeficient mouse strains for PDX engraftment. | NSG (NOD/SCID/IL2Rγnull) offer highest engraftment rates due to lack of adaptive immunity. |
| CellTiter-Glo 3D Assay | Luminescent ATP quantification for 3D cell viability. | Gold-standard for high-throughput PDO drug screens; optimized for spheroids. |
| Cryostor CS10 | Serum-free, defined cryopreservation medium. | Superior recovery of PDX cells and PDOs post-thaw compared to traditional DMSO/FBS mixes. |
Conclusion
Patient-derived xenografts and organoids represent a paradigm shift in cancer modeling, moving research closer to the clinical reality of patient tumors. PDXs offer an unparalleled in vivo context for studying systemic disease and complex microenvironments, while PDOs provide a scalable, manipulable platform for high-throughput discovery and personalized therapeutic guidance. Their complementary nature means the most robust research programs will strategically employ both. The future lies in further standardizing protocols, integrating these models with '-omics' technologies and AI-driven analysis, and strengthening their role in co-clinical trials. By mastering these advanced tools, researchers can de-risk drug development, unravel resistance mechanisms, and ultimately accelerate the delivery of effective, personalized cancer therapies to patients.